Fig. 3.
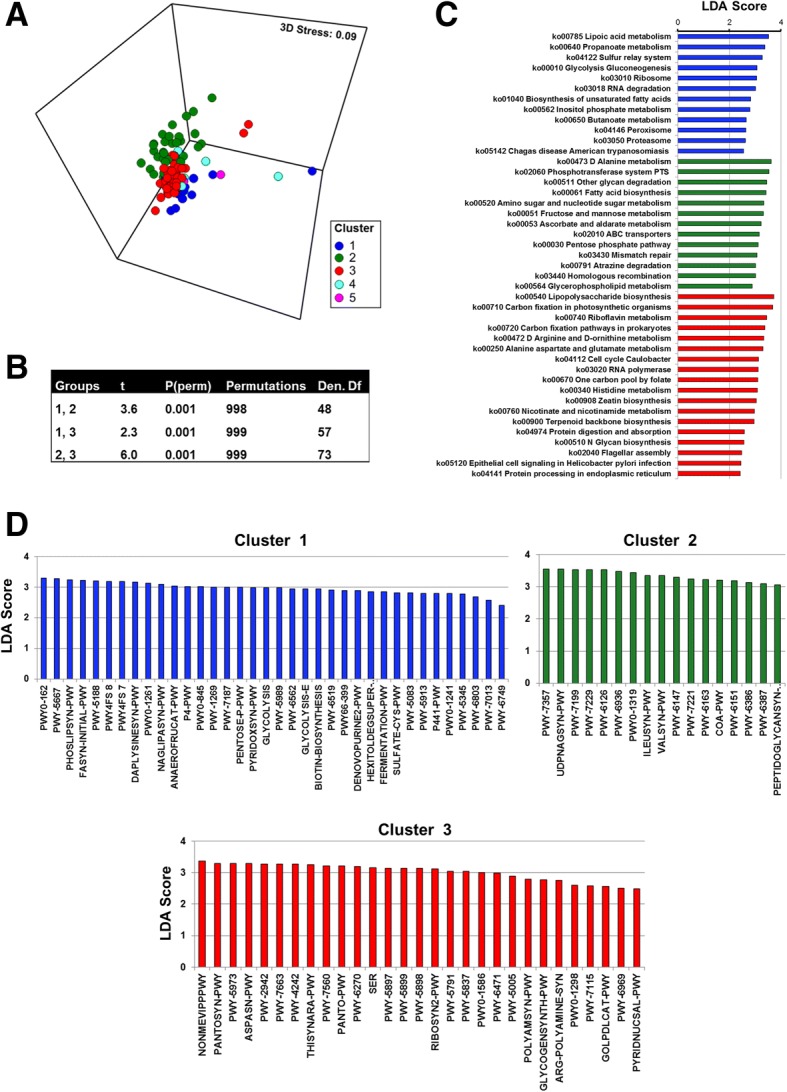
Esophageal community types are functionally distinct. a nMDS plot of Bray–Curtis resemblance generated from square root-transformed KEGG pathway (level 3) relative abundances (generated using HUMAnN2). Clusters from HCA of shotgun data (MetaPhlan2) were overlayed onto the nMDS plot. All available samples were utilized in this analysis. b PERMANOVA across shotgun HCA clusters of Bray–Curtis resemblance generated from square root-transformed KEGG pathway relative abundances. ANOSIM at KEGG pathway level 3 generated a sample statistic of 0.46 and P = 0.001. c KEGG pathways identified using LEfSe analysis to be differentially abundant across each community type. All available samples within each cluster were utilized in this analysis. Blue, cluster 1; green, cluster 2; red, cluster 3. d MetaCyc pathways identified using LEfSe analysis to be differentially abundant across each community type. A full list of pathway names can be found in Additional file 1.14. All available samples within each cluster were utilized in this analysis. ANOSIM for MetaCyc pathways generated a sample statistic of 0.41 and P = 0.001. Blue, cluster 1; green, cluster 2; red, cluster 3
