Fig. 1.
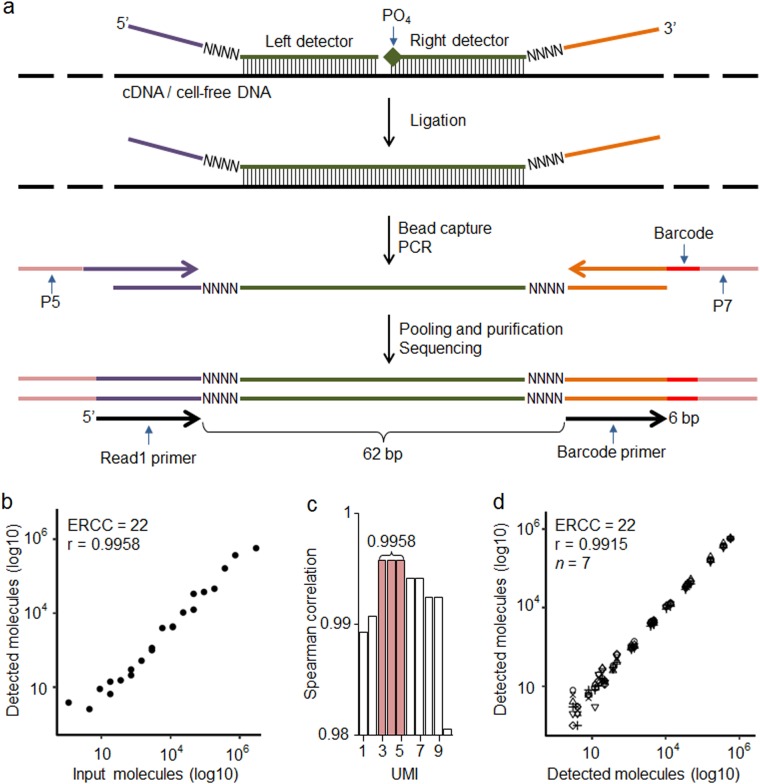
Principle and technical parameters of TAC-seq. a Schematic diagram of the assay to detect specific mRNA or cell-free DNA. Target-specific DNA oligonucleotide detector probes hybridize under stringent conditions to the studied cDNA or cfDNA. Both detector oligonucleotides consist of a specific 27-bp region (green), 4-bp unique molecular identifier (UMI) motif (NNNN), and universal sequences (purple and orange). The right detector oligonucleotide is 5′ phosphorylated. After rigorous hybridization, the pair of detector probes is ligated using a thermostable ligase under stringent conditions. Next, the ligated detectors complexed with the target region are captured with magnetic beads and PCR amplified to introduce sample-specific barcodes and other common motifs that are required for single-read NGS. b Spearman correlation analysis of the input and detected ERCC synthetic spike-in mRNA molecules at UMI threshold 4 (UMI = 4). UMI threshold is defined as the number of detected unique UMI sequences. For example, UMI = 4 indicates that a certain UMI motif is detected at least four times. UMIs are valuable only if the number of UMI combinations (8-bp UMI provides 65,536 variants, for example) is substantially larger than the sum of the target molecules in the studied sample. c Bar plot of Spearman’s correlation analysis of the ERCC input and detected molecules at different UMI thresholds. d Reproducibility of seven technical ERCC replicates (seven different icons on plot) of 22 spike-in molecules at UMI = 4