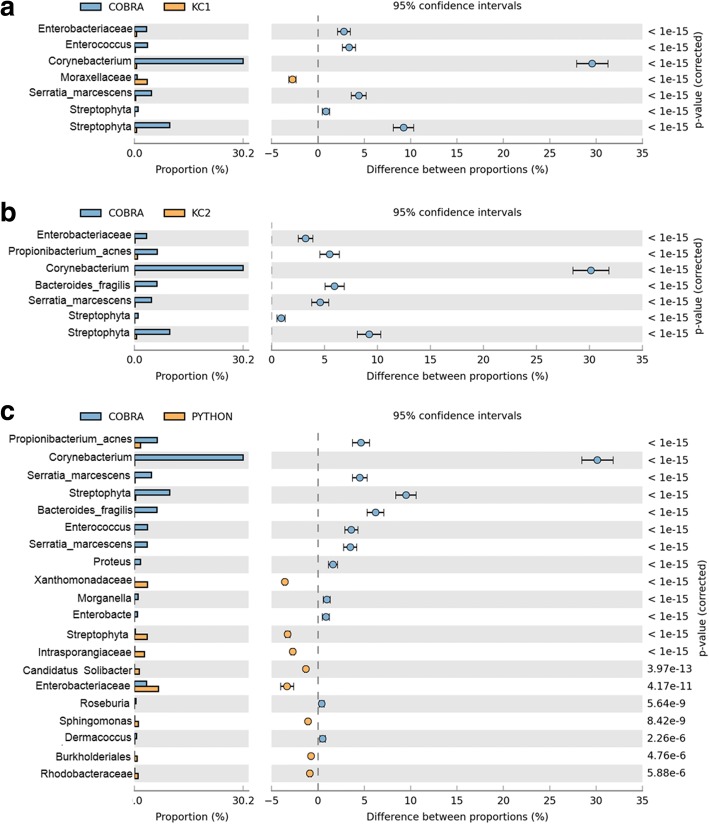
Fig. 5.
Metagenomic profile comparisons of Python, Cobra and King Cobra oral samples determined using STAMP analysis. The comparison includes highly significant phylum to species level. Corrected P-values were calculated based on Fisher’s exact test method using Storey’s FDR approach. P-values < 0.05 were taken for comparison. The bar plot indicated in blue or orange shows a positive or negative difference between read proportions. Differences between samples are shown at 95% confidence intervals a Taxon comparison between Cobra and KC1 samples. Corynebacterium is present in greater abundance in Cobra and lesser abundance in KC1 with positive differences (blue dot), whereas Moraxellaceae is less abundant in Cobra and more abundant in KC1 with negative differences (yellow dot); b Comparison of Cobra and KC2. The most abundant taxon includes Corynebacterium, Bacteroides fragilis and streptophyta, all with positive proportion differences; c Comparison between Cobra and Python samples. Corynebacterium, Propionibacterium acnes and serratia marcescens are highly abundant with positive differences, whereas the species group including Xanthomonadaceae, Streptophyta and Enterobacteriaceae are in greater abundance with negative differences. Here, KC1: King Cobra 1 and KC2: King Cobra 2