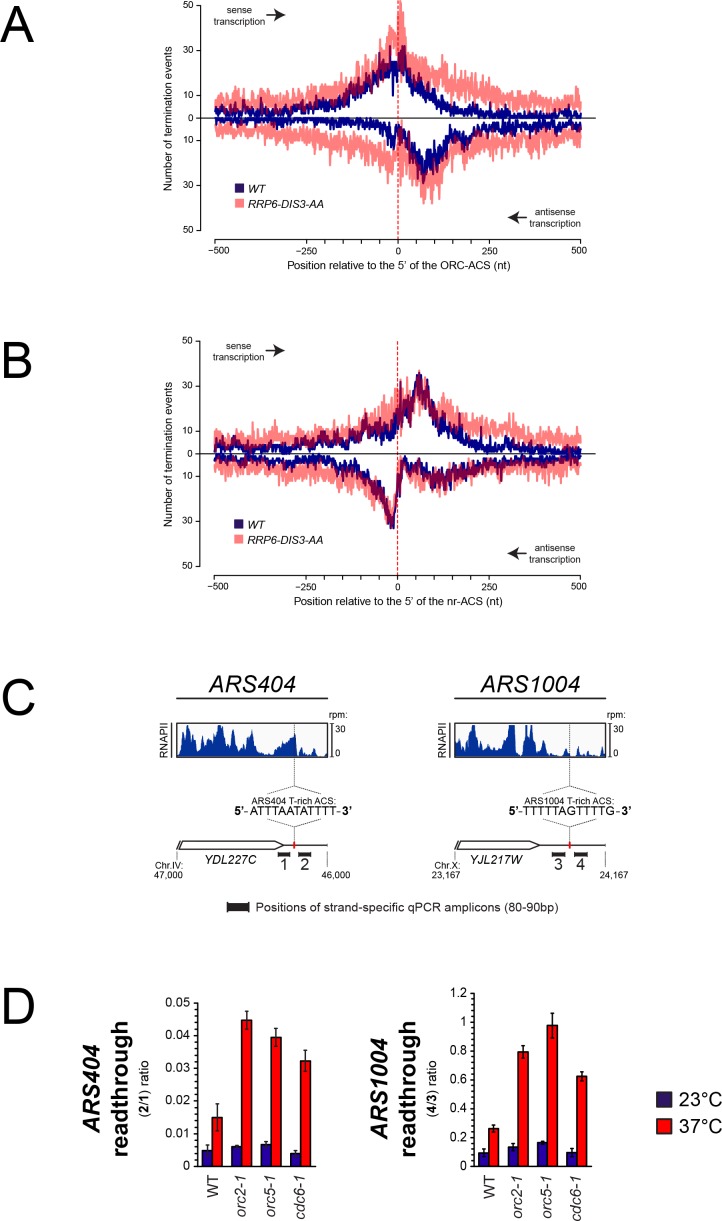
Figure 4. Role of ORC in the roadblock of RNAPII at origins.
(A) Distribution of RNA 3'-ends at genomic regions aligned at ACS sequences recognized by ORC (ORC-ACS) as defined by Eaton et al. (2010) (i.e. defined based on the best match to the consensus associated to each ORC-ChIP peak). Each origin was oriented according to the direction of the T-rich strand of its ORC-ACS and regions were aligned at the 5’ ends of the ORC-ACSs. As in 1B, RNA 3'-ends (Roy et al., 2016) were from transcripts expressed in wild-type cells (blue) or from cells depleted for exosome components (transparent red). At each position around the anchor, presence or absence of an RNA 3'-end was scored independently of the read count. Distributions of RNA 3’-ends both on the sense (top) and the antisense (bottom) strands relative to the ORC-ACSs are plotted. (B). Same as in (A) except that genomic regions were aligned at ACS sequences not recognized by ORC (nr-ACS) as defined by Eaton et al. (2010) (i.e. defined as ACS motifs for which no ORC ChIP signal could be detected). (C). Quantification of the roadblock at individual ARSs. For each ARS, the snapshot includes the upstream gene representing the incoming transcription. The distribution of RNA polymerase II (dark blue) detected by CRAC (Candelli et al., 2018) at ARS404 (left) and ARS1004 (right) oriented according to the direction of their T-rich ACS strands is shown. The positions of the qPCR amplicons used for the RT-qPCR analyses in (D) are indicated. (D). RT-qPCR analysis of transcriptional readthrough at ARS404 and ARS1004. Wild-type, orc2-1, orc5-1 and cdc6-1 cells were cultured at permissive temperature and maintained at permissive (23°C, blue) or non-permissive (37°C, red) temperature for 3 hr. The level of readthrough transcription at ARS404 (left) or ARS1004 ACS (right) was estimated by the ratio of RT-qPCR signals after and before the ACS, as indicated. Data were corrected by measuring the efficiency of qPCR for each couple of primers in each reaction. Values represent the average of at least three independent experiments. Error bars represent standard deviation.