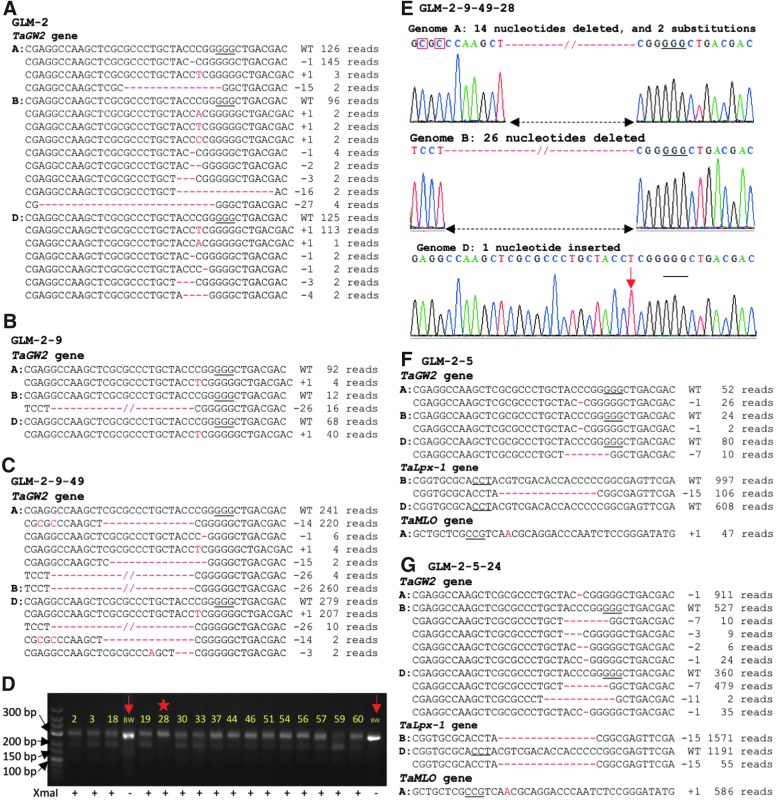
FIG. 4.
Transgenerational CRISPR-Cas9 activity induces new mutations in the TaGW2 and TaLpx-1 genes. NGS reads flanking the GW2T2 target site and their frequencies in (A) T0 line GLM-2, (B) T1 line GLM-2-9, and (C) T2 line GLM-2-9-49 are shown. (D) Restriction enzyme digestion of polymerase chain reaction (PCR) amplicons to screen gw2 knockout mutations in the T3 progenies of line GLM-2-9-49. The GW2T2 flanking region was amplified by PCR and digested with XmaI; non-digested PCR amplicons correspond to mutated GW2T2 target sites. The numbers on the gel image are identifiers of the GLM-2-9-49 progenies. Lanes marked with arrows are PCR products from wild-type plant not digested with XmaI and loaded as controls; the knockout mutant plant was marked with a star. BW, wild-type cultivar Bobwhite. (E) Sanger sequencing of PCR-amplified GW2T2 target sites of T3 line GLM-2-9-49-28. Genome specific primers were used to amplify regions flanking the GW2T2 target sites. Nucleotide substitutions are marked with red rectangles, and the inserted nucleotide is shown by the red arrow. Types and frequencies of mutations at the GW2T2, LPX1T2, and MLOT1 target sites in (F) T1 line GLM-2-5, and (G) T2 line GLM-2-5-24 are shown. WT, wild-type alleles in wheat cultivar Bobwhite; “–” and “+” signs and numbers after them, nucleotides deleted and inserted, respectively. The frequency of each mutation type is shown on the right. The PAM sequences are underlined; the deleted nucleotides are shown with red dashed lines; the insertions and deletions are highlighted in red.