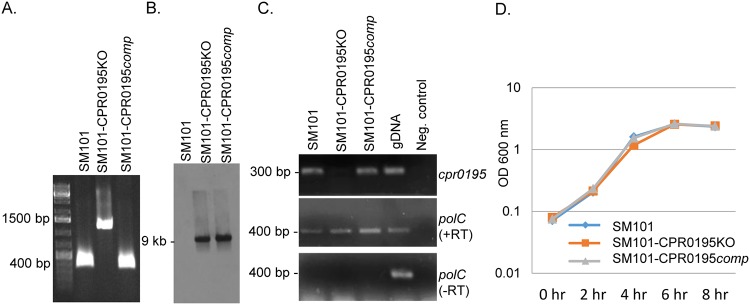
FIG 1.
Characterization of the SM101-CPR0195KO null mutant and SM101-CPR0195comp complementing strain. (A) PCR confirming insertional mutagenesis of the cpr0195 gene in SM101-0195KO. Shown is the cpr0195 PCR product amplified using DNA from wild-type SM101 (lane 2), the SM101-CPR0195KO mutant (lane 3), or the SM101-CPR0195comp complementing strain (lane 4). Note that, compared to the ∼300-bp product amplified using DNA containing a wild-type cpr0195 gene, DNA from the null mutant strain supported amplification of a larger (∼1,200-bp) product due to the insertion of an intron into its cpr0195 gene. (B) Southern blot hybridization of an intron-specific probe with DNA from SM101 (left), SM101-CPR0195KO (middle), or SM101-CPR0195comp (right). DNA from each strain was digested overnight with EcoRI at 37°C and then electrophoresed on a 1% agarose gel. The size of the hybridizing band in the middle and right lanes is shown to the left. Using DNA from wild-type SM101, no intron-specific band was detected, while a single intron-specific band was detected for the SM101-CPR0195KO mutant and complementing strain. (C) RT-PCR analysis for cpr0195 (top panel) or polC (middle panel) transcription in wild-type SM101, the SM101-CPR0195KO mutant, or the complementing strain. SM101 DNA was used as a positive control (gDNA [genomic DNA]). PCRs lacking template DNA acted as a negative control. To show that the RNA preparations from the three strains were free from DNA contamination, these samples were also subjected to PCR without reverse transcription (bottom panel). (D) Growth curves for wild-type SM101, the SM101-CPR0195KO mutant, and the SM101-CPR0195comp strain cultured at 37°C in MDS medium for up to 8 h. Aliquots of each culture were measured every 2 h for their OD600. All experiments were repeated three times, and mean representative values are shown. The markers used in panels A and C were Thermo Fisher 1-kb DNA ladders.