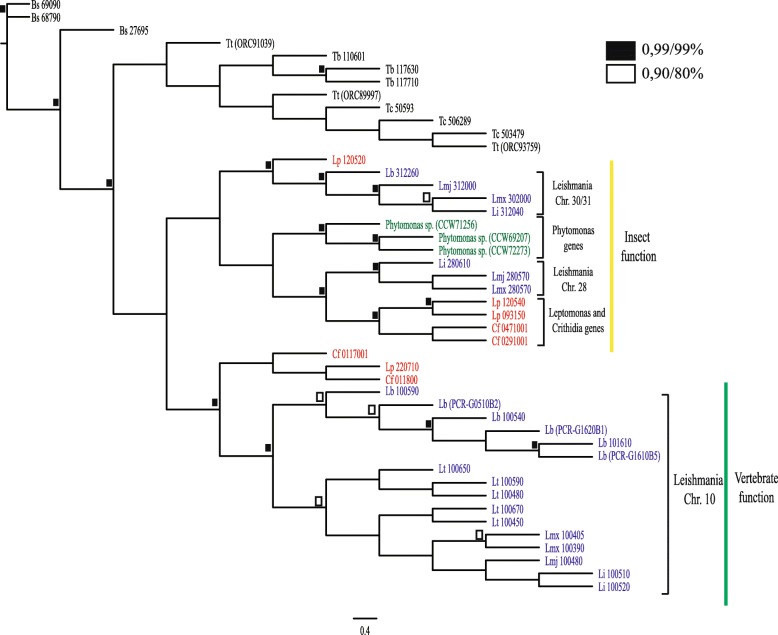
Fig. 3.
Phylogenetic tree comparing selected Leishmania GP63 paralogs with sequences from more distantly related organisms. The tree demonstrates the proximity between the GP63 genes from the Leishmania chromosomes 28 and 31, with GP63 genes from monoxenes trypanosomatids genes, such as Leptomonas pyrrhocoris and Crithidia fasciculata, as well as, genes from a plant tripanosomatid, like Phytomonas sp. Values for highly supported nodes have been replaced by black and white squares, which represents the Bayesian posterior probabilities and bootstrap support for PhyML, respectively. Bs: Bodo saltans, Tt: Trypanosoma theileri, Tb: Trypanosoma brucei, Tc: Trypanosoma cruzi, Lp: Leptomonas pyrrhocoris, Cf: Crithidia fasciculata, Lb: Leishmania braziliensis M2904, Lmj: Leishmania major, Lmx: Leishmania mexicana, Li: Leishmania infantum, Lt: Leishmania tarentolae