FIGURE 3.
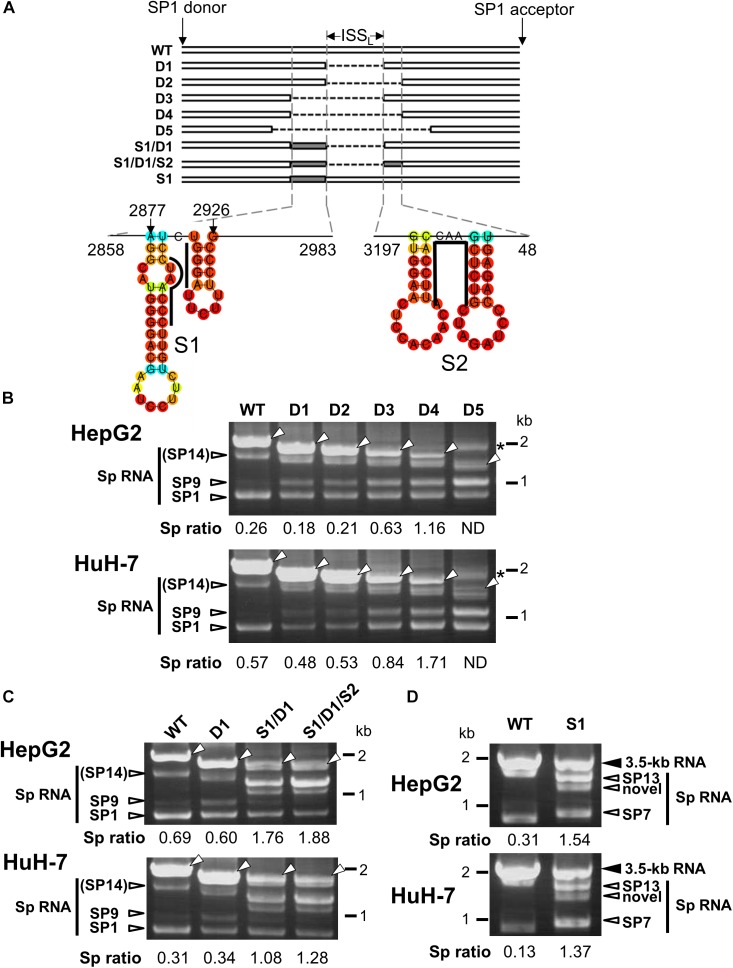
Characterization of the intronic sequence important for silencing of the 3.5 kb RNA splicing. (A) A schematic representation of mutated HBV genomes derived from GT-C with deletions and/or substitutions within the major intron region is shown. Putative secondary structures predicted by the CentroidFold (http://rtools.cbrc.jp/centroidfold/) in nt 2858–2983 and 3197-48 regions are indicated. Each predicted base pair is colored with the heat color gradation from blue to red corresponding to the base-pairing probability. Bold lines are drawn along the sequences targeting to introduce substitution mutations S1 and S2. (B) RT-PCR analysis of HBV RNAs expressed from WT and a series of deletion-mutated HBV genomes (D1–D5) in HepG2 and HuH-7 cells. The quantity ratio of the spliced RNAs to the unspliced RNA (Sp ratio) was determined. ND, not determined. Open arrows, unspliced forms. ∗cDNA products with aberrant sizes. (C) As described in (B) but HBV mutated genomes S1/D1 and S1/D1/S2 in addition to WT and D1 were used for RNA expression. (D) As in (C) but S1 mutant and WT were expressed in the cells. Bands corresponding to unspliced 3.5 kb RNA and its spliced forms (Sp RNA) were indicated.