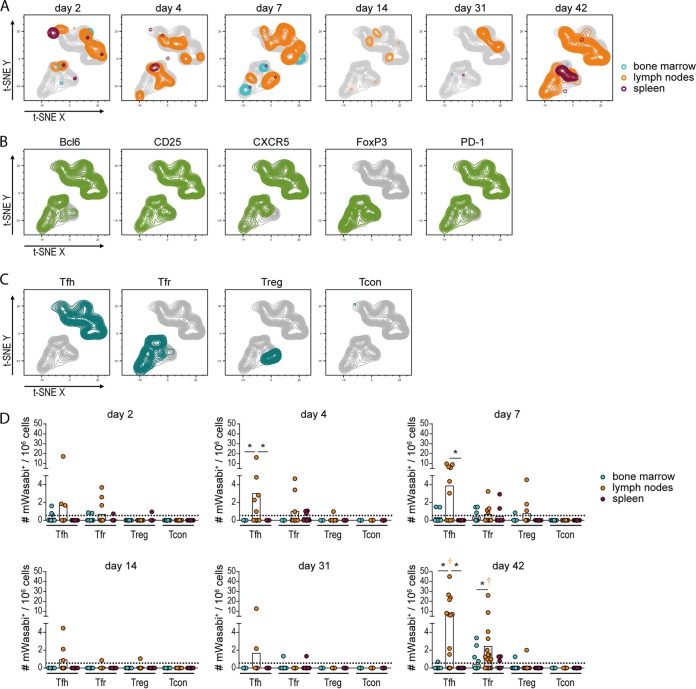
FIG 5.
Analysis of FV-mWasabi-infected CD4+ T cells. C57BL/6 mice were infected with 20,000 SFFU FV-mWasabi, and infected cells were analyzed in bone marrow, lymph nodes, and spleens on days 2, 4, 7, 14, 31, and 42 after FV-mWasabi infection. Cells were subjected to staining with a CD4+ T cell-specific antibody panel, and mWasabi+ CD3+ CD4+ cells were subjected to a t-SNE analysis. (A) Manual gates on samples from individual organs and days of analysis were overlaid on the t-SNE plot to visualize the samples, and manual gates on individual cell markers (B) or manual gates on specific CD4+ T cell types (C) were overlaid on the t-SNE plot to identify cell types. (D) Manual gating was performed to determine the number of follicular helper T cells (Tfh), follicular regulatory T cells (Tfr), regulatory T cells (Treg), or conventional, nonfollicular nonregulatory CD4+ T cells (Tcon). Each circle represents the value for an individual mouse, and bars indicate mean values of groups of mice. The dotted lines indicate the detection limit. The data for each time point were obtained from two (day 4, day 14, and day 31), three (day 2 and day 7), or four (day 42) independent experiments with n = 12 (day 2), 9 (day 4), 10 (day 7), 8 (day 14), 9 (day 31), or 21 (day 42). Statistically significant differences (P < 0.05) between the numbers of infected cells of the different organs within a cell subset are indicated by lines and asterisks; color-coded dagger symbols indicate dominant subsets with significantly higher numbers of FV-mWasabi-infected cells compared to two other subsets within one organ.