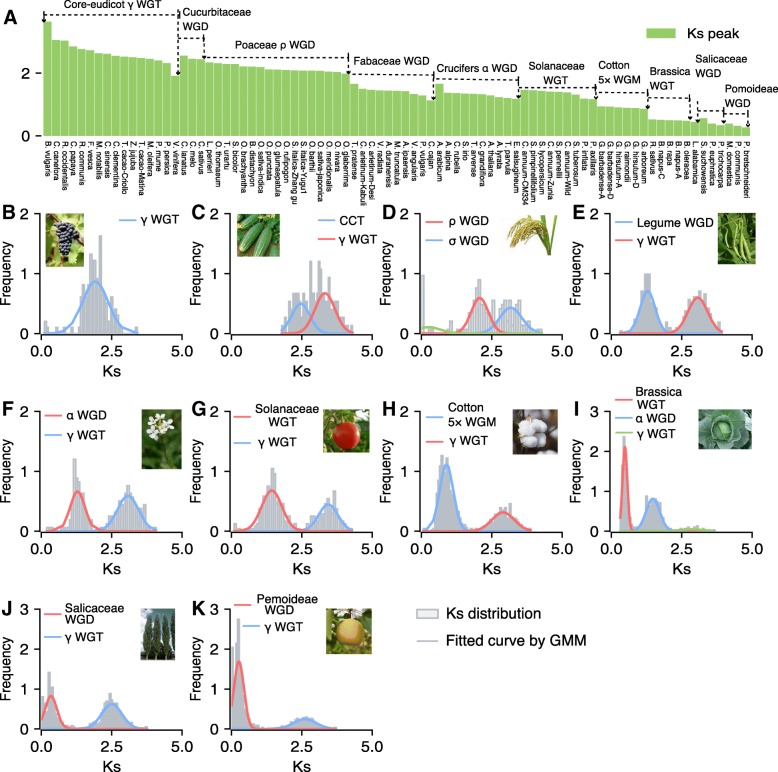
Fig. 2.
Lineage-specific genome duplication events. a The genome duplication events identified in different lineages. A range of Ks values for estimates of individual genome duplication events from different taxa. The Ks distribution in each species was fitted using Gaussian mixture models (GMM). The Ks peaks corresponding to core eudicot γ WGT, cucurbit-common tetraploidization (CCT), Poaceae ρ WGD, Fabaceae common WGD, Brassicaceae α WGD, Solanaceae WGT, cotton 5× WGM, Brassica WGT, Salicaceae WGD, and Pomoideae WGD were respectively detected in Vitis vinifera (b), Cucumis sativus (c), Oryza sativa (d), Phaseolus vulgaris (e), Arabidopsis thaliana (f), Solanum lycopersicum (g), Gossypium raimondii (h), Brassica oleracea (i), Populus trichocarpa (j), and Pyrus bretschneideri (k)