Fig. 3.
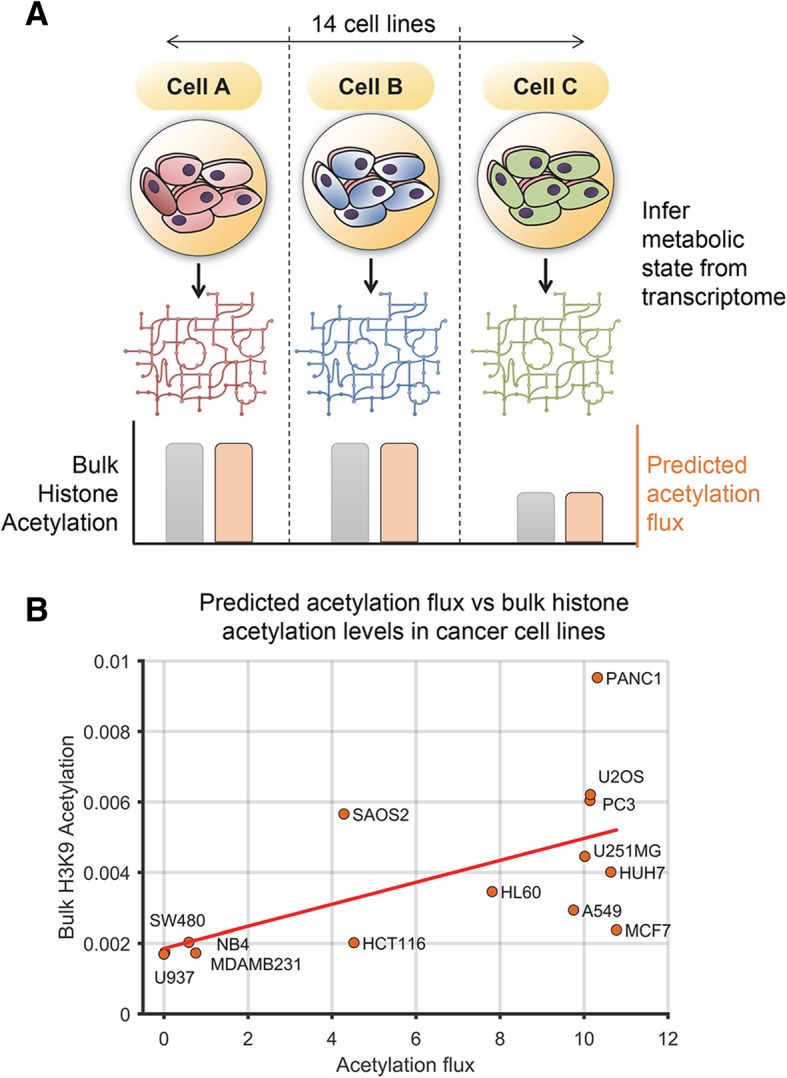
Predicting the effect of the basal metabolic state on histone acetylation levels using FBA. a Schematic of the approach to infer the basal metabolic state using transcriptomic data. The iMAT approach was used to integrate transcriptome data from a panel of cancer cell lines with the human metabolic network model to infer cell-line-specific metabolic state and acetylation flux. b The scatter plot shows the predicted acetylation flux (mmol/gDW cells/h) in each cell line and the bulk histone H3K9 acetylation levels from proteomics. The acetylation flux correlated significantly with the level of bulk histone H3K9 acetylation in these cell lines. The acetylation flux in the high flux group of cell lines is constrained by the availability of carbon units
