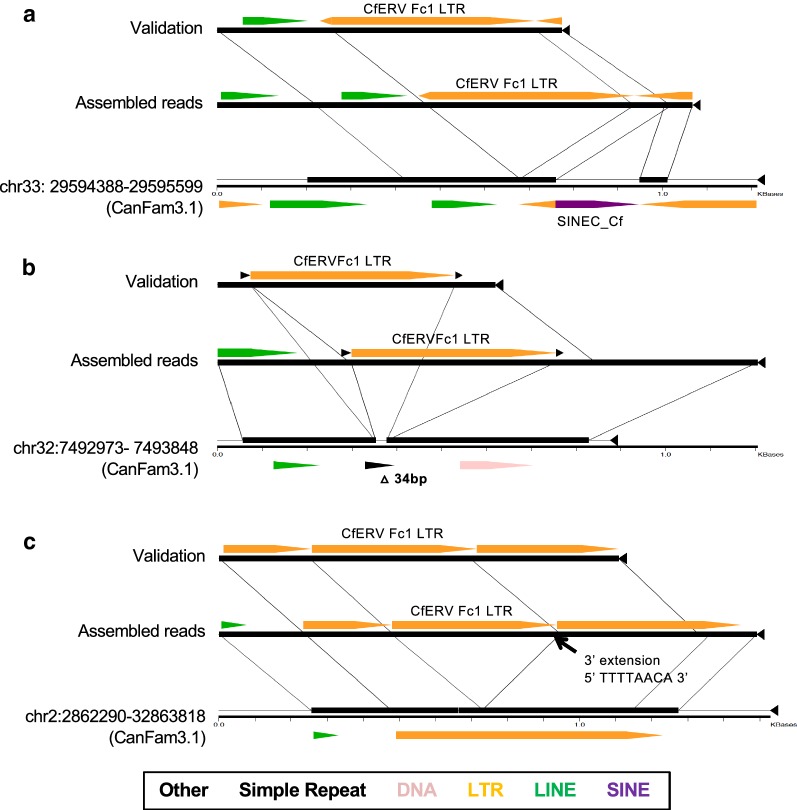
Fig. 4.
Assessment of assembled non-reference alleles. LTR insertions associated with structural variation as captured in assembled Illumina read data. Local three-way alignments were generated for each assembled locus using the program Miropeats [92]. Each consisted of the LTR allele obtained by read assembly, the validated LTR allele obtained by Sanger sequencing of the locus in one individual, and the empty locus as present within the CanFam3.1 reference. Alignments are shown for three representative LTR assemblies. The allele type is labeled at left in each alignment; lines are used to indicate the breakpoint position of the insertion and shared sequence between alleles. a An LTR assembly that includes captured deletion of a bimorphic SINE_Cf insertion present in the CanFam3.1 reference. b An assembled LTR associated with a short 34 bp deletion of sequence that is present in the reference. c A validated assembly of an LTR that included an 8 bp extension relative to the canonical CfERVF1 repeat