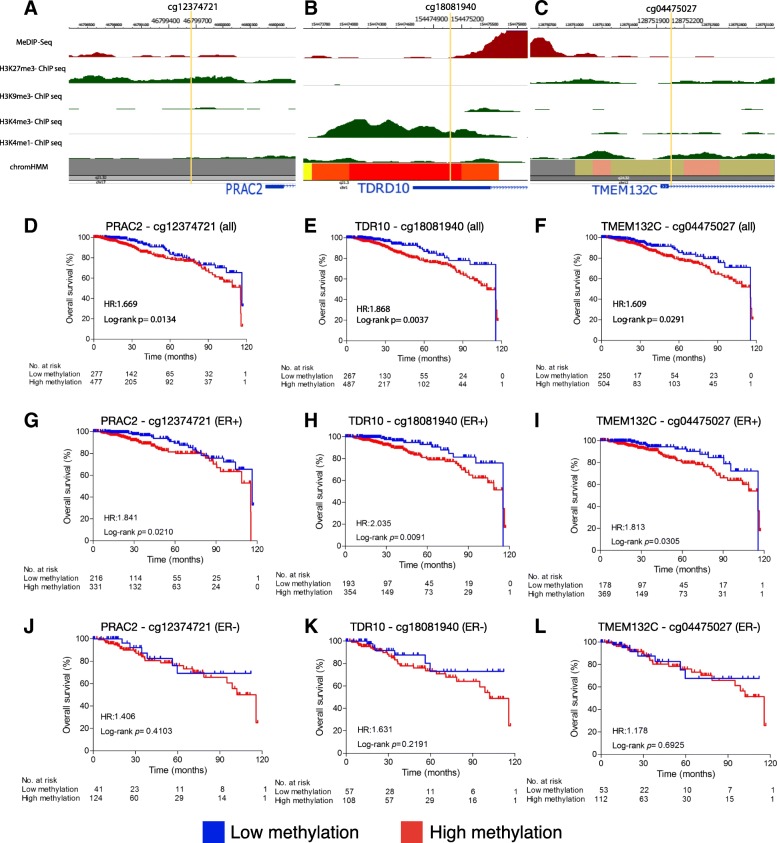
Fig. 4.
Epigenetic analysis of CpGs sites from PRAC2, TDRD10 and TMEM132C (“bottom 7” genes). a MeDIP-Seq data shows that cg12374721 (PRAC2) is hypomethylated in normal breast cells. ChIP-Seq data shows enrichment of H3K27me3 histone repressive marks (green peaks) and lack of H3K4me1 and H3K4me3 active histone marks. ChromHMM classified this region as a poly comb repressive region (grey color). b cg18081940 (TDRD10) and c cg04475027 (TMEM132C) sites are hypomethylated in normal cells and overlap with open chromatin and H3K4me1 and H3K4me3 histone modification peaks associated with active transcription (green peaks). ChromHMM classified b cg18081940 (TDRD10) region as an active TSS (red) and c cg04475027 (TMEM132C) as a bivalent enhancer (dark yellow). d-l Kaplan-Meier curves for the CpG probes located in d PRAC2, e TDRD10 and f TMEM132C showed that hypomethylation is associated with better overall survival. Hypomethylation of the 3 CpGs was also associated with better prognosis in ER-positive samples (g-i), but not in ER-negative samples (j-l). Based on the AUC, a cut-off value was established for each probe in order to distinguish hypomethylated patients (blue) from hypermethylated patients (red). The following cut-offs were used: d 0.5503, corresponding to the 37th percentile (PRAC2-cg12374721); e 0.5243, 35th percentile (TDRD10-cg18081940); f 0.4014, 33rd percentile (TMEM132C-cg04475027); g 0.5503, 39th percentile; h 0.5243, 35th percentile; i 0.4014, 33rd percentile; j 0.5503, 25th percentile; k 0.5243, 35th percentile; l 0.4014, 32th percentile