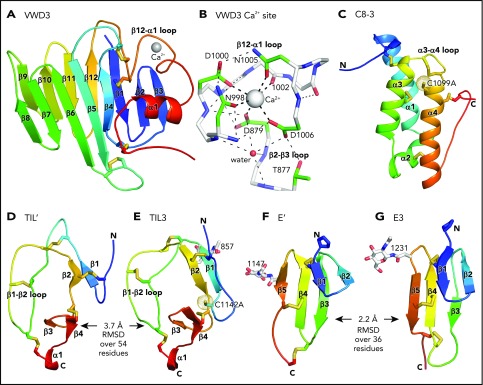
Figure 3.
Structure of D assembly modules. (A,C-G) Each module is shown rainbow colored to trace the path the polypeptide takes to create the fold, from N terminus (blue) to C terminus (red). (B) The Ca2+-binding site. Mainchain carbons are shown in white, with sidechain carbons in green. Ca2+ and a water oxygen are shown as spheres. Coordination bonds to the Ca2+ and hydrogen bonds are shown as thick and thin dashed lines, respectively. Other representations are as in Figure 1. Superposition of TIL domains required the Deep Align server27; E modules were superimposed with PyMol. Superimposed structures are shown separated horizontally on the page. Structural differences between modules are shown as root mean square deviation of Cα atoms.