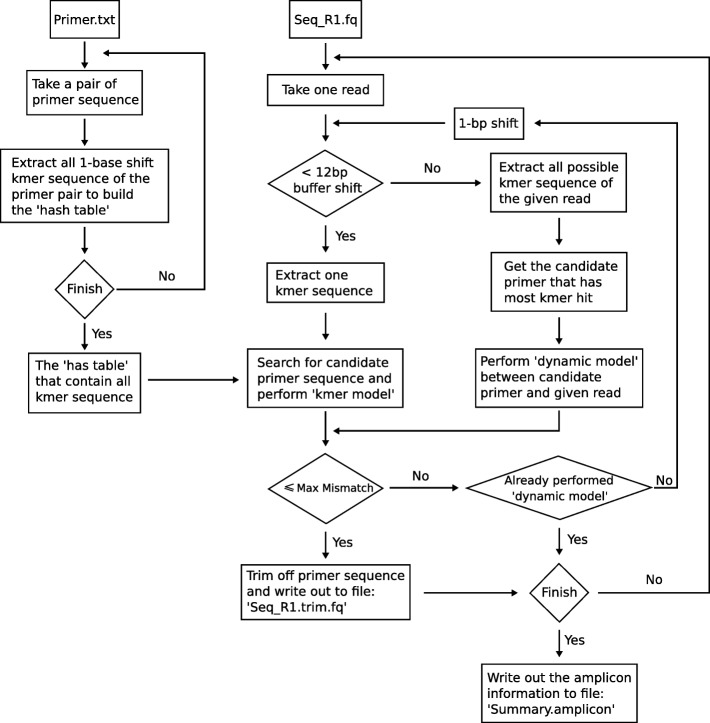
Fig. 1.
Workflow of pTrimmer. The flowchart shows the details of the program. The program takes a primer sequence file and the two FASTQ files (paired-end sequencing) or one (single-end sequencing) as input. The primer sequence file contains three necessary fields for the forward primer, reverse primer and insert length of amplicon. pTrimmer will first extract all possible 1-base shift kmer sequence of both forward primer and reverse primer to build the ‘hash table’. Kmer sequence will also be extracted from FASTQ read with 1-bp shift each time to search for best candidate primer from ‘hash table’. The 12-bp buffer shift length enable the program has 12 times to find the best primer sequence with ‘k-mers model’. If fails, the program will automatically switch to ‘dynamic model’. The ‘dynamic model’ could extract all possible 1-bp shift kmer sequence and get the best candidate primer that has most kmer hit. Then the program will perform dynamic programing between candidate primer sequence and given read