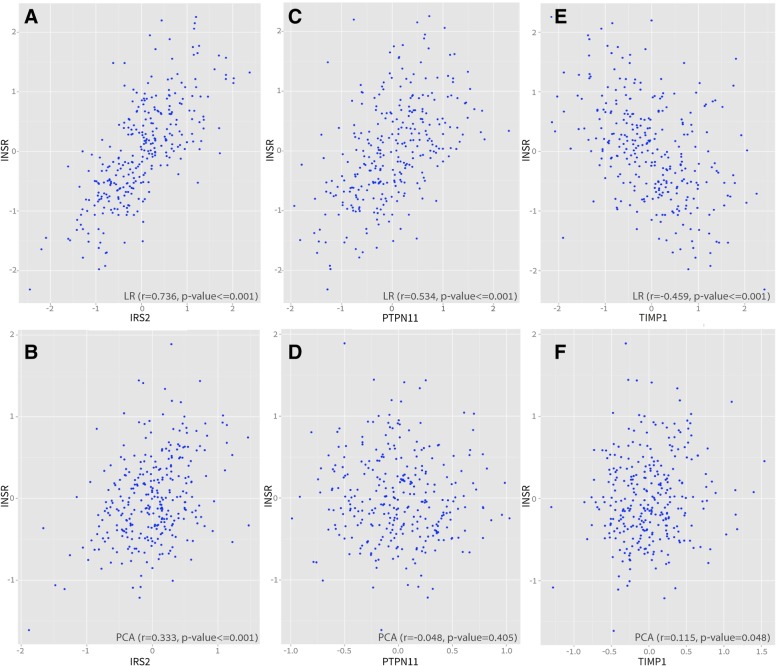
Fig. 3.
Examples of Spearman correlation coefficients of three true gene-gene associations. These are calculated following LR-based adjustment (using the linear regression model for known confounders) and principal components-based adjustment of the GTEx Adipose Subcutaneous dataset. The example genes are derived from the insulin signaling mechanism. a, b Example of co-expression plot of LR and PCA-based adjustment for INSR-IRS2 association. c, d Example of co-expression plot of LR and PCA-based adjustment for INSR-PTPN11 association. e, f Example of co-expression plot of LR and PCA-based adjustment for INSR-TIMP1 association. The y-axis presents the INSR (insulin receptor) expression for each sample (depicted by circles) and the x-axis the expression values of the relevant gene. Spearman correlation coefficients and p-values are presented at the top of each plot. It can be seen that removing most of the variability using principal components-based adjustment (PCA) can result in eliminating a desired biological signal, e.g., PTPN11 has a significant correlation coefficient to INSR (r = 0.534, p-value < 0.001) following the LR adjustment and non-significant (r = ~ 0, p-value = 0.405) following the PCA adjustment