Figure 2.
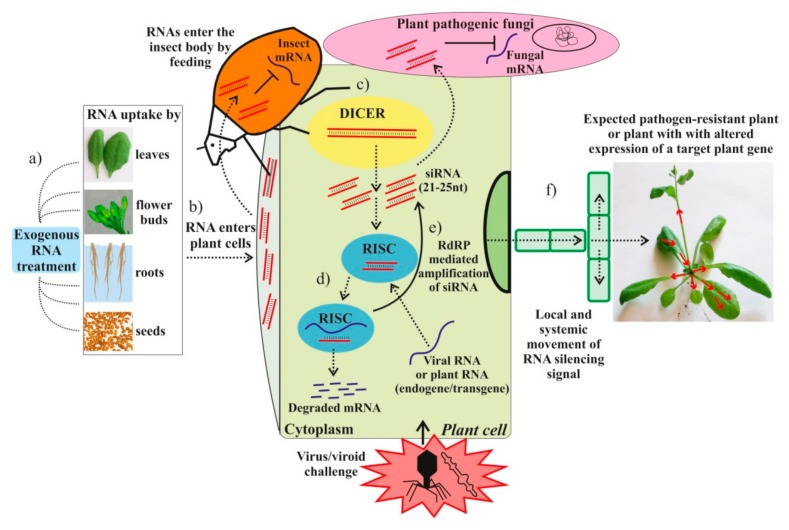
Schematic representation of exogenous RNA applications for RNA interference (RNAi) induction and degradation of the target plant pathogen or endogenous plant mRNAs. (a) Exogenous artificial RNA provided in a solution and applied onto plant leaves, flower buds, roots, or seeds. (b) The exogenous RNAs are taken up and transported into the cytoplasm via an undefined mechanism. (c) The dsRNA or hpRNA molecules are recognized by a ribonuclease, DICER-like (DICER), which cleaves the dsRNA into siRNAs. (d) The siRNAs are then incorporated in the RNA-induced silencing complex (RISC) that guides sequence-specific degradation or translational repression of homologous mRNAs. (e) The components of the siRNA/mRNA complex can be amplified into secondary siRNAs by the action of RNA-dependent RNA-polymerase (RdRP). (f) Movement of the RNA silencing signal between plant cells and through the vasculature. Dashed arrows depict different steps of the RNAi induction process and dsRNA/siRNA movement between plant cells and plant pathogens. The solid arrow depicts the RdRP-mediated amplification of siRNA. Red arrows depict the local and systemic movement of the RNA silencing signal in the plant.