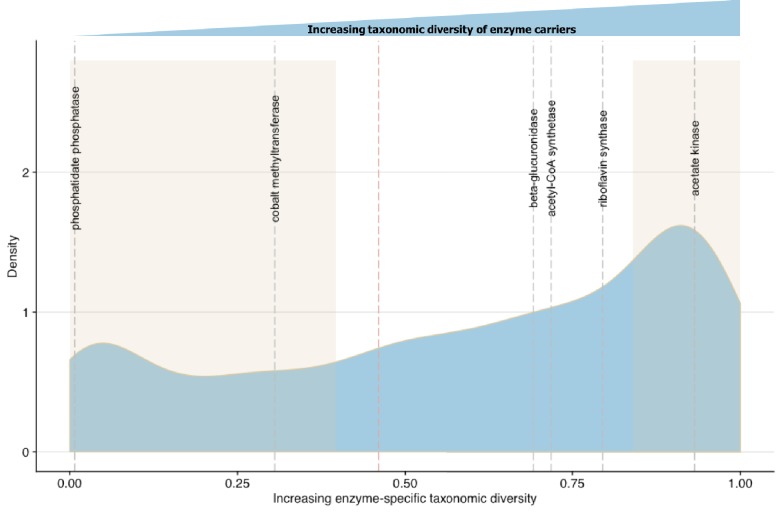
Figure 7. Linking enzymatic functions with taxonomic diversity.
The Simpson index was adapted to describe enzyme-specific taxonomic dominance and diversity based on enzyme abundance in taxonomy-linked gene counts across healthy individuals in the Integrative Human Microbiome Project (Proctor et al., 2014). We define a microbial enzyme as high dominance and low taxonomic diversity if its Simpson index value falls below 0.46 (red dotted line), the mean value across all enzymes. Dominance-diversity values for gut microbiota functions that fall above or below the mean are highlighted by gray dashed lines and include the following enzymes and pathways: phosphatidate phosphatase (0.007353), cobalt-precorrin-2 C(20)-methyltransferase (0.305502) from the Vitamin B12 synthesis pathway, β-glucuronidase (0.691618), Acetyl-CoA synthase (0.718163) which is involved in the production of propionate from complex carbohydrates, riboflavin synthase (0.794781) from the riboflavin synthesis pathway and acetate kinase (0.931892) which is involved in acetate production. The shaded regions indicate the range of EDsD values that are one standard deviation above and below the mean and reflect the most broadly distributed functions and most specialized functions.