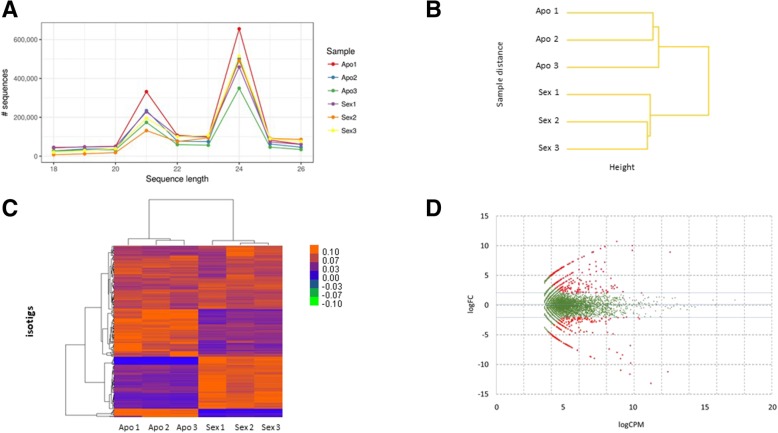
Fig. 2.
General analysis of sRNA sequencing outputs derived from sexual and apomictic P. notatum floral samples. a Length distribution of small RNAs sequences after trimming and adapter removal. b Clustering of the number of small RNA sequences aligned to a Paspalum floral transcriptome reference obtained from the three replicates of sexual and apomictic samples. c Heat map of the 250 transcripts carrying the largest number of small RNA reads. Orange and blue colours indicate higher and lower numbers of sRNAs mapping in each transcript, respectively. d Mean-difference plot for differential small RNA transcripts coverage in sexual and apomictic floral tissues. Only transcripts carrying n > 5.0 were considered. Green dots: non-significant differential expression. Red dots: significant differential expression (logFC >|2|, FDR < 0.05). Positive logFC: upregulation in apomictic samples. Negative logFC: upregulation in sexual samples