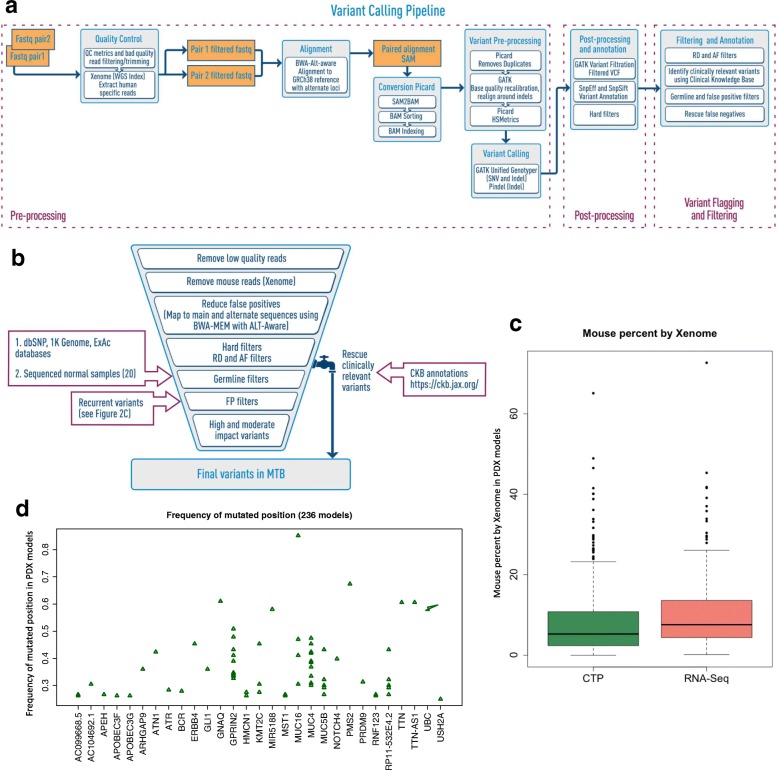
Fig. 2.
A multi-step variant filtering and “rescue” strategy to accurately identify somatic mutations in PDX tumors. a Overview of the PDX variant calling workflow for engrafted tumors in the absence of paired-normal tumor samples and in the presence of mouse stroma. (see Methods for details). b Overview of the filter and rescue strategy used in the variant calling workflow for JAX PDXs. MTB: Mouse Tumor Biology Database, RD: Read depth, AF: Allele-frequency, FP: False positives, CKB: JAX Clinical Knowledgebase. c Proportion of mouse reads classified by Xenome for CTP and RNA sequencing data across all PDX models. d The recurrent frequencies of the mutated positions (after germline filtering) for various genes that were found to be recurrent in more than 25% of PDX samples. These were identified as additional false positive variants due to sequencing errors or mapping issues