Abstract
Bacterial diseases of swine are a kind of multifactorial and uncontrollable diseases that commonly exist in pig farms all over the world and will lead to huge economic losses every year. In this study, a detailed and overall survey was carried out to better understand the prevalence and antimicrobial susceptibilities of bacterial diseases from 2013 to 2017 in China. A total of 19673 bacterial strains were isolated from 44175 samples collected from 9661 pig farms that distributed in 16 Chinese major pig breeding provinces. The results showed that the average isolation rates of Streptococcus suis (SS), Haemophilus parasuis (HPS), Escherichia coli (E. coli), Pasteurella multocida (Pm), Actinobacillus pleuropneumoniae (APP), Brodetella bronchiseptica (Bb), Salmonella enteria (SE), Erysipelothrix rhusiopathiae (E. rhusiopathiae) were 16.9%, 9.7%, 6.3%, 3.4%, 0.3%, 1.5%, 2.3% and 0.9%, respectively. The isolate rates of E. coli, APP and SE showed an increasing trend from 2013 to 2017. The seasonal prevalence characteristics of SS, HPS and Pm were obviously higher from April to August for first two bacteria and higher at February, March, April, and October for Pm. The dominant serotypes for SS, HPS were serotype 2 and serotype 5 (changed from serotype 4), respectively. The SS, HPS, and Pm showed very high antibiotic resistance rates to almost 8 common antibiotics (β-lactam, aminoglycoside, macrolides, lincomycin, tetracycline, quinolone, polymyxin, and sulfonamide) and an obvious increasing trend of antibiotic resistance rates from 2013 to 2017. In conclusion, the study provides detailed information on the prevalence and antimicrobial susceptibilities of different bacterial pathogens of swine from 2013 to 2017 in China. These data can provide a foundation for monitoring epidemiological patterns of bacterial diseases in the Chinese swine herds, as well as provide insight into potential antibiotic resistance profiles in these pathogens.
Subject terms: Cellular microbiology, Bacterial pathogenesis
Introduction
Bacterial diseases heavily affect the health of swine, especially, respiratory system and digestive system diseases which are reported to be associated with intensive pig production1. For example, Streptococcus suis (SS) can lead to septicemia, meningitis, arthritis, acute death and even affect the health of human2. Haemophilus parasuis (HPS) and Actinobacillus pleuropneumoniae (APP) can lead to dyspnea, pneumonia, pleurisy, and progressive emaciation3. Escherichia coli (E. coli) and Salmonella enteria (SE) can lead to diarrhea and other gastrointestinal diseases4,5. Pasteurella multocida (Pm) and Brodetella bronchiseptica (Bb) can together lead to swine atrophic rhinitis6. Meanwhile, Erysipelothrix rhusiopathiae (E. rhusiopathiae) can lead to cutaneous necrosis, endocarditis, and arthritis7. In total, they are all the reasons that heavily affect the health of swine and lead to economic losses of pig industry.
Because of their complexity and indeterminacy, bacterial diseases are very difficult to control. Among them, serotype is a very important factor, we know that bacteria often have many kinds of serotypes and they often lack cross-protection between different serotypes. Furthermore, serotype has obvious distribution difference between different regions. For example, the main serotypes of SS in Canada are serotype 2, 1/2, 3 and 48. However, serotype 2, 9, 7 SS are dominant in China. Therefore, it is very important to know the main serotypes of common bacterial pathogens for controlling bacterial diseases and it will also supply guidance for developing vaccines.
Antimicrobial agents are widely used in bacterial diseases of swine, especially, respiratory tract infections and diarrheal diseases. Antibiotics have been widely used in livestock industries since the early 1990s in China. The average usage of veterinary antibiotics has reached approximately 6000 tons annually9 and most of them are used as feed additives, such as tetracyclines, sulfonamides, fluoroquinolones, macrolides and others10. The widely use of veterinary antibiotics greatly contributes to the development of livestock industries. However, there are also some problems existed in usage of antibiotics, such as abusing of antibiotics in fodder, drinking water, and injection and violating withdrawal time, which all will lead to an increase in antibiotic resistance rates10,11. In addition, it not only affects the development of the pig industry but also threats the health of human12. Therefore, how to improve the production efficiently and decrease the use of antibiotics have become urgent and difficult problems.
Though significant progress has been made in the last few decades in reducing the prevalence of bacterial diseases, there is still an increasing concern over the losses associated with diseases. So, the isolation rates, regional and seasonal distribution, serotype survey, and antimicrobial susceptibilities of major bacteria were analyzed to understand, prevent and control bacterial diseases in China.
Materials and Methods
Samples collection
All pig tissue samples were collected from 16 Chinese major pig breeding provinces from 2013 to 2017 and delivered to the Animal Disease Diagnostic Center of Huazhong Agricultural University for identification of bacterial pathogens, which is a reference lab in China where people send pig samples for diagnosis (Fig. 1). All samples that were sent to our reference lab with a total of 44175 samples from 9661 pig farms were collected and chosen for bacterial isolation and identification (Table 1). The pig farms had a wide variety of management types and herd size with small-, medium- and large-scale commercial pig farms as well as various types of backyard farms. The tissues came from suspected sick pigs and were transported to our reference lab. The collected samples included lung, heart, spleen, joint fluid, intestine, brain, liver, trachea, effusion and so on (Fig. 2). Then, sterile operation had been taken to avoid cross contamination and all samples were processed for bacterial isolation immediately. The research was approved by the Ethics Committee of the Faculty of Veterinary Medicine of the Huazhong Agricultural University. All procedures regarding the animal care and testing were carried out according to the recommendation of Hubei provincial public service facilities.
Figure 1.
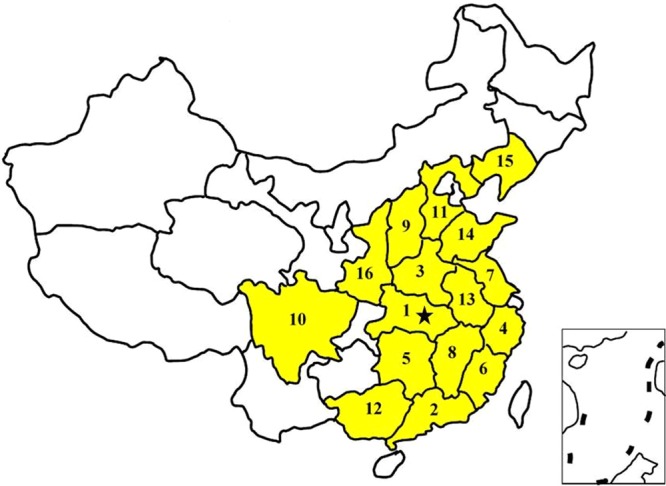
Geographic distribution of different collected samples. The position of asterisk is the location of our diagnostic laboratory. 1–16 represented different provinces where samples were collected and the ranking of the number of samples. 1: Hubei (12897, 29.2%), 2: Guangdong (12389, 28.0%), 3: Henan (5890, 13.3%), 4: Zhejiang (3902, 8.8%), 5: Hunan (3080, 7.0%), 6: Fujian (960, 2.2%), 7: Jiangsu (897, 2.0%), 8: Jiangxi (853, 1.9%), 9: Shanxi (696, 1.6%), 10: Sichuan (652, 1.5%), 11: Hebei (533, 1.2%), 12: Guangxi (295, 0.7%), 13: Anhui (263, 0.6%), 14: Shandong (257, 0.6%), 15: Liaoning (157, 0.4%), 16: Shanxi (119, 0.3%) and others (332, 0.8%).
Table 1.
Samples information.
| Years | No. pig farms | No. samples | No. isolated bacteria |
|---|---|---|---|
| 2013 | 1374 | 7496 | 3116 |
| 2014 | 2989 | 9452 | 3926 |
| 2015 | 2974 | 9320 | 4495 |
| 2016 | 1051 | 8226 | 3543 |
| 2017 | 1273 | 9681 | 4593 |
| Total | 9661 | 44175 | 19673 |
Figure 2.
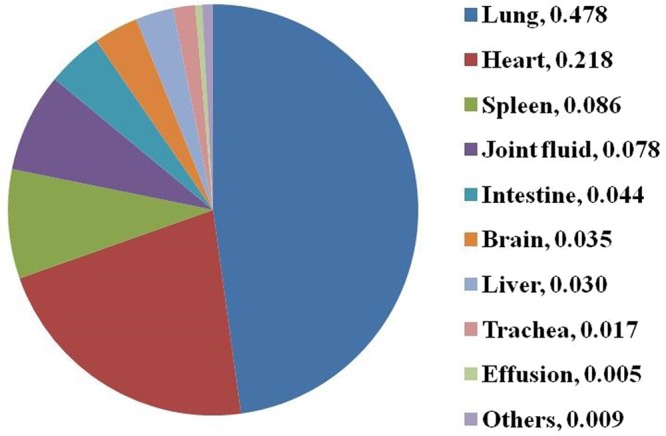
Tissue sources of collected samples. Tissue sources of these samples were lung (21127, 47.8%), heart (9612, 21.8%), spleen (3801, 8.6%), joint fluid (3445, 7.8%), intestine (1942, 4.4%), brain (1556, 3.5%), liver (1314, 3.0%), trachea (765, 1.7%), effusion (235, 0.5%) and others (378, 0.9%).
Culture conditions, primers, and identification methods
Common bacterial pathogens were isolated and identified from Chinese pig farms, such as SS, HPS, Pm, E. coli, APP, Bb, SE, E. rhusiopathiae, and others. Tryptic Soy Broth (TSB), Tryptic Soy Agar (TSA) (Difco Laboratories, Detroit, USA), Shigella and Salmonella Agar and MacConkey Agar (HuanKai Microbial, Guangdong, China) medium were used and 10 μg/mL of nicotinamide adenine dinucleotide (NAD) and 5% (v/v) inactivated cattle serum (Zhejiang Tianhang Biotechnology, Zhejiang, China) were added if necessary. Primers used for identification and serotyping were listed in Table 2 and Supplementary Table 1, respectively.
Table 2.
Primers used in this study.
| Strains | Gene | Name | Sequence (5′ → 3′) | Amplicons size (bp) | References |
|---|---|---|---|---|---|
| Streptococcus suis | gdh |
JP4 JP5 |
GCA GCGTATTCTGTCAAACG CCATGGACA GATAAA GATGG |
689 | 47 |
| Haemophilus parasuis | 16S rRNA |
HPS-1 HPS-2 |
GGCTTCGTCACCCTCTGT GTGATGAGGAAGGGTGGTGT |
822 | 48 |
| Pasteurella multocida | kmt1 |
KMT1T7 KMT1SP6 |
ATCCGCTATTTACCCAGTGG GCTGTAAACGAACTCGCCAC |
457 | 49 |
| Escherichia coli | uidA |
Ec-1 Ec-2 |
AAAACGGCAAGAAAAAGCAG GCGTGGTTACAGTCTTGCG |
147 | 50 |
| Actinobacillus pleuropneumoniae | apxIVA |
APXIVA-1 APXIVA-2 |
TGGCACTGACGGTGATGA GGCCATCGACTCAACCAT |
442 | 51 |
| Brodetella bronchiseptica | fla |
Fla4 Fla2 |
TGGCGCCTGCCCTATC AGGCTCCCAAGAGAGAAA |
237 | 52 |
| Salmonella enteria | invA |
INVA-1 INVA-2 |
ACAGTGCTCGTTTACGACCTGAAT AGACGACTGGTACTGATCGATAAT |
580 | 53 |
| Erysipelothrix rhusiopathiae | ER |
ER1 ER2 |
CGATTATATTCTTAGCACGCAACG TGCTTGTGTTGTGATTTCTTGACG |
937 | 54 |
| Universal primer | 16S rRNA |
27F 1492R |
AGAGTTTGATCCTGGCTCAG TACGGCTACCTTGTTACGACTT |
1466 | 21 |
All plates were incubated at 37 °C for 24 to 48 h. After this isolation stage, the strains were identified by colony morphology, Gram-staining characteristics and oxidase (Gram-negative bacilli) or catalase tests. Phenotypic methods or standard biochemical procedures were used to identify suspected bacteria of SS, HPS, Pm, E. coli, APP, Bb, SE, E. rhusiopathiae, and others based on the previous studies4,13–19. To further confirm the phenotypic and biochemical results, PCR methods of specific genes were used based on the references mentioned in Table 2. When PCR identification results were not consistent with the results of phenotypic or standard biochemical procedures, the strains were further identified by 16S rRNA sequencing20,21 (Table 2). All isolated bacteria were freeze-dried and kept at −80 °C.
Serotype identification of SS and HPS
Parts of isolated SS (273/1149, 231/1469, 168/1719, 136/1357 and 128/1796 strains from 2013 to 2017, respectively) and HPS (331/868, 225/883, 206/1231, 179/570 and 154/711 strains from 2013 to 2017, respectively) strains were randomly chosen to do serotype identification based on previously described methods22,23. According to the reports, SS can be divided into 33 serotypes based on the difference of capsular polysaccharide, and HPS can be divided into 15 serotypes based on the difference of capsular loci22,23.
Antimicrobial susceptibility test
Antimicrobial susceptibility test was performed based on the standard Clinical and Laboratory Standards Institute (CLSI) guidelines for susceptibility testing. Disk diffusion (DD) was performed according to CLSI M2 A12 Ed. 12 (2015) standards. Tests were performed according to the manufacturer instructions. Briefly, a sterile cotton-tipped swab was dipped into the bacterial suspension (0.5 McFarland) and streaked in three directions across TSA agar plates containing 10 μg/mL of NAD and 5% (v/v) inactivated cattle serum. The plates were dried for 2–3 min, then disks were placed on the agar surface, which contained common antibiotics that used in Chinese pig farms, such as cephradine (CE, 30 μg), ceftriaxone (CRO, 30 μg), amoxicillin (AML, 10 μg), ampicillin (AMP, 10 μg), streptomycin (S, 10 μg), gentamicin (CN, 10 μg), spectinomycin (SH, 100 μg), kanamycin (K, 30 μg), amikacin (AK, 30 μg), neomycin (N, 30 μg), spiramycin (SP, 100 μg), azithromycin (AZM, 15 μg), lincomycin (MY, 2 μg), clindamycin (DA, 2 μg), doxycycline (DO, 30 μg), ofloxacin (OFX, 5 μg), ciprofloxacin (CIP, 5 μg), enrofloxacin (ENR, 5 μg), polymyxin B (PB, 300 μg) and trimethoprim (W, 1.25 μg) (Oxoid, UK). The plates were incubated for 2 days in an aerobic atmosphere at 37 °C. At the end of the incubation period, the diameters of the zones of growth inhibition were measured and the final reference standard based on CLSI24,25.
Statistical analysis
Statistical analyses were undertaken with SAS version 9.0 (SAS Institute Inc.). Univariate association between variables and isolation rates of different bacteria were determined by using univariate ordinary logistic regression analysis and Chi square test. P < 0.05 was considered to be significant.
Results
Sample sources of bacterial pathogens
44175 tissues samples from 9661 pig farms were collected and 19673 bacterial pathogens were isolated from 2013 to 2017 (Table 1). The samples were collected from Hubei (12897, 29.2%), Guangdong (12389, 28.0%), Henan (5890, 13.3%), Zhejiang (3902, 8.8%), Hunan (3080, 7.0%), Fujian (960, 2.2%), Jiangsu (897, 2.0%), Jiangxi (853, 1.9%), Shanxi (696, 1.6%), Sichuan (652, 1.5%), Hebei (533, 1.2%), Guangxi (295, 0.7%), Anhui (263, 0.6%), Shandong (257, 0.6%), Liaoning (157, 0.4%), Shanxi (119, 0.3%) and others (332, 0.8%) (Fig. 1), respectively. Among them, Hubei and Guangdong provinces are the major places of sample sources, which contained over half of them (57.2%, 25287/44175). On the one hand, the tissue sources of these samples were lung (21127, 47.8%), heart (9612, 21.8%), spleen (3801, 8.6%), joint fluid (3445, 7.8%), intestine (1942, 4.4%), brain (1556, 3.5%), liver (1314, 3.0%), trachea (765, 1.7%), effusion (235, 0.5%) and others (378, 0.9%) (Fig. 2), respectively.
The proportion of isolated bacterial pathogens
The isolation rates of different bacterial pathogens were presented at Fig. 3. From the results, we knew that 8 kinds of common bacteria (SS, HPS, E. coli, Pm, APP, Bb, SE, and E. rhusiopathiae) were isolated from pigs and the top three kinds of bacteria were SS, HPS and E. coli, which contained about 73.9% of isolated bacteria.
Figure 3.
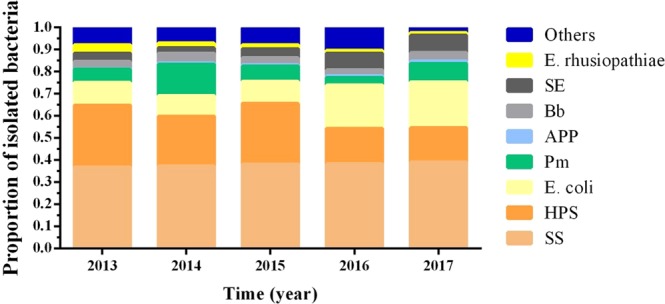
The proportion of isolated bacterial pathogens from 2013 to 2017. Different bacteria were labeled using different colors and the height of the column represented the isolation rates. The successfully isolated bacteria were SS (1149, 36.9%), HPS (868 27.9%), E. coli (320, 10.3%), Pm (188, 6.0%), APP (9, 0.3%), Bb (101, 3.2%), SE (117, 3.8%), E. rhusiopathiae (117, 3.8%), and others (247, 7.9%) in 2013, SS (1469, 37.4%), HPS (883, 22.5%), E. coli (367, 9.3%), Pm (556, 14.2%), APP (23, 0.6%), Bb (173, 4.4%), SE (94, 2.4%), E. rhusiopathiae (85, 2.2%), and others (276, 7.0%) in 2014, SS (1719, 38.2%), HPS (1231, 27.4%), E. coli (451, 10.0%), Pm (309, 6.9%), APP (27, 0.6%), Bb (143, 3.2%), SE (185, 4.1%), E. rhusiopathiae (82, 1.8%), and others (348, 7.7%) in 2015, SS (1357, 38.3%), HPS (570, 16.1%), E. coli (694, 19.6%), Pm (120, 3.4%), APP (28, 0.8%), Bb (91, 2.6%), SE (269, 7.6%), E. rhusiopathiae (46, 1.3%), and others (368, 10.4%) in 2016, SS (1796, 39.1%), HPS (711, 15.5%), E. coli (955, 20.8%), Pm (379, 8.3%), APP (51, 1.1%), Bb (176, 3.8%), SE (362, 7.9%), E. rhusiopathiae (55, 1.2%), and others (108, 2.4%) in 2017.
The prevalence characteristics of different bacterial pathogens
Based on the isolation rates of different bacterial pathogens, the prevalence characteristic of single-pathogen was analyzed (Fig. 4). The detail prevalence trends of SS (Fig. 4A), HPS (Fig. 4B), E. coli (Fig. 4C), Pm (Fig. 4D), APP (Fig. 4E), Bb (Fig. 4F), SE (Fig. 4G), E. rhusiopathiae (Fig. 4H) were shown in Fig. 4. From the results, we knew that the prevalence of SS (from 15.3% to 18.6%), E. coli (from 3.9% to 9.9%), APP (from 0.1% to 0.5%) and SE (from 1.0% to 3.7%) increased from 2013 to 2017. However, the prevalence of HPS (from 13.2% to 6.9%) and E. rhusiopathiae (from 1.6% to 0.6%) declined from 2013 to 2017. Different with other bacterial pathogens, Pm and Bb showed no significant prevalence trends from 2013 to 2017. But Pm and Bb exhibited a similar changing trend among different years.
Figure 4.
The isolation rates of different bacterial pathogens from 2013 to 2017. (A–H) represented the isolation rates of SS (A), HPS (B), E. coli (C), Pm (D), APP (E), Bb (F), SE (G), E. rhusiopathiae (H), respectively. Error bars represented the standard deviation of isolation rates of 12 months.
The seasonal prevalence characteristics of SS, HPS, and Pm
To investigate the seasonal prevalence characteristics of SS, HPS, and Pm, the number of samples and isolation rates of every month were calculated when samples were collected from 2013 to 2017. Results showed that the rates of isolation across the whole year of SS ranged from 13.6% to 23.5% with the highest isolation rate recorded in June (Fig. 4). While the monthly isolation rates of HPS ranged from 7.7% to 11.7% with the highest isolation rate recorded in July. By comparing the seasonal prevalence characteristics of SS and HPS, a similar phenomenon was found that they all had higher isolation rates from April to August. However, Pm showed higher isolation rates in February, March, April, and October ranging from 2.6% to 5.2% within a whole year (Fig. 5).
Figure 5.

The seasonal prevalence characteristics of SS, HPS, and Pm. The accurate isolation rates of all bacterial pathogens in the different months from 1–12 were 13.6% (428/3151), 15.0% (388/2586), 14.9% (626/4207), 20.4% (836/4097), 19.2% (764/3992), 23.5% (899/3826), 18.9% (725/3843), 19.5% (697/3571), 13.6% (501/3673), 15.5% (517/3337), 14.1% (620/4389) and 13.9% (488/3503), respectively. As a whole, the differences in the isolation rates of SS, HPS, and Pm in 12 months were all significant (p < 0.05). Statistical analysis was performed using the χ2 test.
The serotype of SS and HPS
To better understand the most common bacterial pathogens in Chinese pig farms, a part of isolated SS and HPS were chosen to do serotyping experiments every year. The results showed that the major SS existed in Chinese pig farms were serotype 2, 9, 7, 3, 1, 5 from 2013 to 2017 and the highest isolated rate of SS was still serotype 2, but it declined sharply from 2013 (52.4%) to 2017 (33.6%). However, there were several other serotypes of SS increased over time. Such as serotype 7 (from 7.3% to 19.5%) and serotype 9 (from 12.1% to 25.8%) (Fig. 6A). Meanwhile, the major HPS existed in Chinese pig farms were serotype 4, 5, 13, 2, 1, 14, 10 and 12 from 2013 to 2017. Nevertheless, the proportion of different serotype of HPS changed obviously and the biggest change of HPS was serotype 4 (from 30.8% to 20.8%) and 5 (from 12.1% to 31.8%) which were the most popular serotypes of HPS in 2013 and 2017, respectively. At the same time, the proportion of HPS serotype 1 (from 7.9% to 1.3%), 2 (from 9.1% to 5.0%) and 10 (from 4.4% to 1.3%) declined, serotype 13 (from 10.9% to 17.5%) increased and serotype 12 and 14 did not show any change trends (Fig. 6B).
Figure 6.
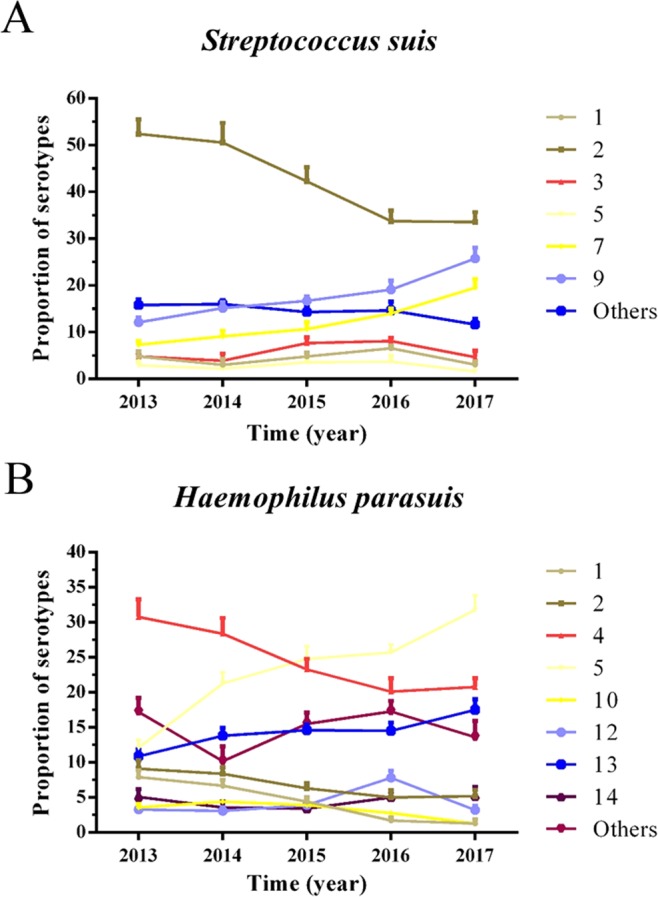
The proportion of different serotypes of SS (A) and HPS (B). The number of SS strains chosen for serotyping included 273, 231, 168, 136, and 128 strains from 2013 to 2017, respectively. Meanwhile, the number of HPS strains included 331, 225, 206, 179, and 154 strains from 2013 to 2017, respectively. As a whole, the differences in serotype 2 and 7 of SS from 2013 to 2017 were significant (p < 0.05), and the differences in serotype 4, 5 and 13 of HPS from 2013 to 2017 were significant (p < 0.05). Statistical analysis was performed using the χ2 test.
Antibiotic resistance rates of SS, HPS, and Pm
A total of 8 kinds of antibiotics have been chosen to do antimicrobial susceptibility tests including β-lactam (cephradine, ceftriaxone, amoxicillin and ampicillin), aminoglycoside (streptomycin, gentamicin, spectinomycin, kanamycin, amikacin and neomycin), macrolides (spiramycin and azithromycin), lincomycin (lincomycin and clindamycin), tetracycline (doxycycline), quinolone (ofloxacin, ciprofloxacin and enrofloxacin), polymyxin (polymyxin B) and sulfonamide (trimethoprim) antibiotics. Antibiotic resistance rates of SS, HPS, and Pm were presented in Table 3. Results showed that antibiotic resistance rates of SS to aminoglycoside, macrolides, lincomycin, tetracycline polymyxin, and sulfonamide antibiotics were all over 60% and it exhibited increasing antibiotic resistance rates in β-lactam (from 11.6% to 19.7%) and quinolone (from 34.7% to 49.9%) antibiotics (Fig. 7A). Meanwhile, the antibiotic resistance rates of HPS and Pm to different antibiotics are obviously lower than SS. But, HPS showed significant increasing antibiotic resistance rates to all kinds of antibiotics except sulfonamide which showed a strong antibiotic resistance rate to HPS (Fig. 7B). Just like HPS, Pm also had increasing antibiotic resistance rates to all kinds of antibiotics except lincomycin and quinolone antibiotics (Fig. 7C).
Table 3.
Antibiotic resistance rates of SS, HPS, and Pm.
| Bacteria/Antibiotic resistance rates | SS | HPS | Pm | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2013 (95) | 2014 (136) | 2015 (182) | 2016 (188) | 2017 (217) | 2013 (62) | 2014 (89) | 2015 (124) | 2016 (83) | 2017 (90) | 2013 (29) | 2014 (42) | 2015 (43) | 2016 (18) | 2017 (55) | |
| CE | 0.063 | 0.059 | 0.055 | 0.112 | 0.129 | 0.000 | 0.011 | 0.065 | 0.048 | 0.044 | 0.000 | 0.000 | 0.047 | 0.056 | 0.036 |
| CRO | 0.189 | 0.162 | 0.143 | 0.207 | 0.217 | 0.129 | 0.045 | 0.113 | 0.193 | 0.167 | 0.310 | 0.310 | 0.256 | 0.389 | 0.400 |
| AML | 0.074 | 0.110 | 0.126 | 0.138 | 0.189 | 0.081 | 0.135 | 0.169 | 0.205 | 0.189 | 0.103 | 0.143 | 0.140 | 0.167 | 0.218 |
| AMP | 0.137 | 0.147 | 0.192 | 0.324 | 0.253 | 0.145 | 0.146 | 0.185 | 0.325 | 0.556 | 0.069 | 0.095 | 0.070 | 0.278 | 0.291 |
| S | 0.800 | 0.772 | 0.665 | 0.888 | 0.866 | 0.226 | 0.449 | 0.500 | 0.639 | 0.544 | 0.241 | 0.190 | 0.209 | 0.333 | 0.436 |
| CN | 0.832 | 0.941 | 0.907 | 0.830 | 0.788 | 0.161 | 0.213 | 0.274 | 0.446 | 0.333 | 0.207 | 0.571 | 0.558 | 0.611 | 0.509 |
| SH | 0.463 | 0.632 | 0.577 | 0.670 | 0.544 | 0.065 | 0.112 | 0.169 | 0.470 | 0.300 | 0.103 | 0.143 | 0.140 | 0.278 | 0.255 |
| K | 0.905 | 0.934 | 0.802 | 0.872 | 0.853 | 0.194 | 0.258 | 0.194 | 0.349 | 0.311 | 0.069 | 0.262 | 0.279 | 0.556 | 0.527 |
| AK | 0.979 | 0.978 | 0.951 | 0.840 | 0.903 | 0.306 | 0.427 | 0.363 | 0.542 | 0.422 | 0.552 | 0.524 | 0.512 | 0.778 | 0.636 |
| N | 0.989 | 0.971 | 0.940 | 0.979 | 0.839 | 0.306 | 0.382 | 0.331 | 0.892 | 0.744 | 0.690 | 0.667 | 0.581 | 0.778 | 0.873 |
| SP | 0.874 | 0.912 | 0.901 | 0.979 | 0.995 | 0.548 | 0.483 | 0.419 | 0.904 | 0.989 | 0.759 | 0.810 | 0.721 | 0.944 | 0.982 |
| AZM | 0.832 | 0.816 | 0.709 | 0.872 | 0.903 | 0.065 | 0.056 | 0.113 | 0.193 | 0.156 | 0.000 | 0.071 | 0.070 | 0.056 | 0.055 |
| MY | 0.989 | 0.993 | 0.962 | 0.851 | 0.935 | 0.274 | 0.213 | 0.210 | 0.470 | 0.744 | 0.897 | 0.905 | 0.884 | 0.778 | 0.836 |
| DA | 0.958 | 0.971 | 0.945 | 0.814 | 0.986 | 0.452 | 0.438 | 0.427 | 0.482 | 0.989 | 0.966 | 0.952 | 0.953 | 0.778 | 0.909 |
| DO | 0.989 | 0.993 | 0.973 | 0.926 | 0.862 | 0.113 | 0.090 | 0.113 | 0.325 | 0.267 | 0.276 | 0.286 | 0.302 | 0.611 | 0.636 |
| OFX | 0.326 | 0.360 | 0.385 | 0.335 | 0.341 | 0.081 | 0.124 | 0.210 | 0.217 | 0.556 | 0.034 | 0.071 | 0.116 | 0.056 | 0.109 |
| CIP | 0.421 | 0.441 | 0.429 | 0.473 | 0.770 | 0.403 | 0.348 | 0.274 | 0.651 | 0.533 | 0.172 | 0.143 | 0.186 | 0.111 | 0.109 |
| ENR | 0.295 | 0.279 | 0.346 | 0.410 | 0.387 | 0.145 | 0.157 | 0.234 | 0.289 | 0.378 | 0.069 | 0.095 | 0.093 | 0.111 | 0.164 |
| PB | 0.937 | 0.912 | 0.896 | 0.867 | 0.687 | 0.065 | 0.034 | 0.040 | 0.229 | 0.544 | 0.034 | 0.071 | 0.116 | 0.167 | 0.218 |
| W | 0.726 | 0.824 | 0.615 | 0.697 | 0.429 | 0.774 | 0.708 | 0.621 | 0.880 | 0.767 | 0.448 | 0.595 | 0.651 | 0.889 | 0.655 |
CE: Cephradine; CRO: Ceftriaxone; AML: Amoxycillin; AMP: Ampicillin; S: Streptomycin; CN: Gentamicin; SH: Spectinomycin; K: Kanamycin; AK: Amikacin; N: Neomycin; SP: Spiramycin; AZM: Azithromycin; MY: Lincomycin; DA: Clindamycin; DO: Doxycycline; OFX: Ofloxacin; CIP: Ciprofloxacin; ENR: Enrofloxacin; PB: Polymyxin B; W: Trimethoprim.
Figure 7.
Antibiotic resistance rates of SS (A), HPS (B) and Pm (C) from 2013 to 2017. Antibiotics were chosen for antimicrobial susceptibility tests included β-lactam (cephradine, ceftriaxone, amoxicillin and ampicillin), aminoglycoside (streptomycin, gentamicin, spectinomycin, kanamycin, amikacin and neomycin), macrolides (spiramycin and azithromycin), lincomycin (lincomycin and clindamycin), tetracycline (doxycycline), quinolone (ofloxacin, ciprofloxacin and enrofloxacin), polymyxin (polymyxin B), and sulfonamide (trimethoprim) antibiotics. As a whole, the differences in resistance rates of SS, HPS, and Pm to all antibiotics from 2013 to 2017 were significant (p < 0.05), except resistance rates of SS to β-lactam and Pm to quinolone. Statistical analysis was performed using the χ2 test.
Discussion
Based on samples collection information of geographical distribution, tissue sources and bacterial isolation rates, results had been concluded that SS, HPS, E. coli, Pm, APP, Bb, SE, and E. rhusiopathiae were ubiquitous in almost all Chinese pig farms (Figs 1 and 2). From the previous reports, we know that SS, HPS, Pm, APP, and Bb are the major bacteria which seriously influence the respiratory system of pigs26, E. coli and SE are the major pathogens of digestive system4,5, and E. rhusiopathiae will lead to endocarditis, cutaneous necrosis and arthritis7. They are all important pathogens that influence pig growth and productivity, and will lead to huge economic losses to the pig industry27,28. Though significant progresses have been made in the last few decades in reducing the prevalence of animal diseases, there is still an increasing concern over the economic losses associated with diseases that cause a reduction in production efficiency. Therefore, it is very important to know the prevalence characteristics and geographical distribution of these bacteria to control bacterial diseases efficiently.
This study fully evaluates the occurrence of common bacterial pathogens in Chinese pig farms. The results showed that the proportion of SS was highest among all isolated bacteria, followed by HPS, E. coli, Pm, APP, Bb, SE, and E. rhusiopathiae. Meanwhile, the order of the isolation rates of these bacterial pathogens was the same as the proportion of isolated bacteria (Fig. 3). Compared with the previous reports, higher isolation rates of SS, HPS, E. coli, and Pm were observed from pigs with respiratory diseases in agreement with some previous studies29–31. Whereas, lower isolation rates of Bb, APP, and E. rhusiopathiae were obtained20,29,32 (Fig. 4). On the other hand, although Pm and Bb did not show any obvious epidemic characteristics from 2013 to 2017, the changing trend of their isolation rates was almost the same which may associate with the characteristic that they always exist side by side and they are all the common pathogens of swine atrophic rhinitis33.
By analyzing the isolation rates of SS, HPS, and Pm among different months, the characteristics of their seasonal distribution were shown clearly. The isolation rates of SS and HPS were obviously higher in the warm season (from April to August) than the cold season because hot and wet climate is beneficial to the growth and transmission of SS and HPS34. However, the seasonal distribution of Pm was different with SS and HPS, Pm showed a significantly higher isolation rate from February to April (Fig. 5) which is almost the same with Bb as the report20. The reasons that lead to the phenomenon may be due to the discovery that Pm is always commonly existed with Bb which also has a high isolation rate from January to April20,33. On the other hand, because of cold and wet weather, the ventilation installations are always closed in pig farms to keep warm which contributes to the spread of Pm. Therefore, the climate is an important factor to monitor and control bacterial diseases.
It is reported that SS can be divided into at least 33 kinds of serotypes35 and HPS can also be divided into 15 kinds of serotypes36. However, their pathogenicity is different between different serotypes and they lack of cross-protection37,38. So, it is very important to know the clinical serotypes of SS and HPS for controlling bacterial diseases. Our serotyping results showed that the major serotypes SS existed in China were 2, 9, 7, 3, 1, 5 (Fig. 6A). But, it was different with other reports, especially, the decline of serotypes 3, 4, 8 and 1/2 and the increase of serotypes 7 and 9, which suggested the changing trend of different serotypes of SS in China39. On the other hand, SS of serotype 2 declined sharply from 2013 (52.4%) to 2017 (33.6%) which may be associated with the use of SS vaccine of serotype 2 in Chinese pig farms (Fig. 6A). A similar changing trend also appeared between different serotype of HPS. Based on serotyping results, serovars 4, 5 and 13 were still the major serotype of HPS which were the same as the reports before in China, North America, and Spanish40–42. Nevertheless, the percentage of different serotype of HPS varied largely and the main serotype of HPS changed from serotype 4 in 2013 (30.8%) to serotype 5 in 2017 (31.8%) which were all the main serotype of HPS existed in Chinese pig farms. There was also an obvious change in other serotypes (Fig. 6B). The main reasons that lead to the difference in serotype of SS and HPS may be due to the difference of environment, antibiotic, and the use of bacterial vaccines. Therefore, it is very important to change the serotype of the inactivated vaccines in controlling these diseases.
Because of the good effect of antibiotics in promoting growth and preventing infection, large amounts of antimicrobial agents are still being used in modern swine production around the world. This, in turn, would facilitate the emergence and development of antimicrobial resistance. So, it is very important to monitor antibiotic resistance rates of clinical pathogens43. In the experiments, we analyzed antibiotic resistance rates of SS, HPS, and Pm from 2013 to 2017. The results indicated that SS, HPS, and Pm all showed similar and very high antibiotic resistance rates to 8 kinds of detected antibiotics with the reports15,44,45. Thereinto, the antibiotic resistance rates of SS to these 8 kinds of antibiotics were all over 60% except β-lactam and quinolone. Meanwhile, these pathogens all displayed a rapid increase in antibiotic resistance rates of the common used antibiotics in China from 2013 to 2017, which maybe indicate irregular and excessive use of antibiotics. Based on the previous reports, the ways that some bacteria produce antibiotic resistance to aminoglycoside, macrolides, lincomycin, tetracycline, polymyxin, and sulfonamide mainly due to obtainment of exogenous resistance genes. However, the main reasons that lead to antibiotic resistance to β-lactam and quinolone are because of the mutation of drug targets, which is obviously more difficult than the first one46. Therefore, the difference of resistance mechanism maybe an important reason that the resistance rates of aminoglycoside, macrolides, lincomycin, and sulfonamide is obviously higher than that to β-lactam and quinolone in SS, HPS, and Pm. In total, reasonable application of antibiotics has become an urgent issue for the control of bacterial diseases.
Conclusions
In summary, the identification and analysis results of 44175 collected samples showed that the main bacteria in Chinese pig farms were still SS, HPS, E. coli, and Pm. However, the isolation rates of different strains and serotypes of SS and HPS have obviously changed from 2013 to 2017. For example, SS, E. coli, APP, and SE displayed an increasing isolation rates. Nevertheless, HPS and E. rhusiopathiae displayed reversed results. The main serotypes of HPS had changed from serotype 4 to serotype 5, and the proportion of serotype 2 SS also decreased sharply from 2013 to 2017. Meanwhile, SS and HPS had an higher isolation rate in hot season. But, February, March, April, and October were the main seasons for the isolation of Pm. In addition, Antimicrobial susceptibility test indicated that SS, HPS, and Pm presented very high and increasing resistance rates to 8 kinds of common antibiotics. In conclusion, the study provides us very detailed information on the prevalence and antimicrobial susceptibilities of several main bacteria in China from 2013 to 2017, which help us to understand, prevent and control bacterial diseases of Chinese pig farms.
Supplementary information
Acknowledgements
This research was supported by the China Agriculture Research System (No. CARS-35).
Author Contributions
Conceptualization, Bingzhou Zhang and Qigai He; Data curation, Bingzhou Zhang and Xugang Ku; Formal analysis, Bingzhou Zhang and Xugang Ku; Funding acquisition, Qigai He; Investigation, Bingzhou Zhang, Xugang Ku, Xuexiang Yu, Qi Sun, Hao Wu, Long Guo and Xibiao Tang; Methodology, Bingzhou Zhang, Xugang Ku, Fangzhou Chen and Qigai He; Project administration, Qigai He; Software, Bingzhou Zhang, Fangzhou Chen and Xiaoqian Zhang; Supervision, Qigai He; Visualization, Bingzhou Zhang, Xiaoqian Zhang, Long Guo and Xibiao Tang; Writing – original draft, Bingzhou Zhang; Writing – review & editing, Bingzhou Zhang Fangzhou Chen and Qigai He.
Competing Interests
The authors declare no competing interests.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1038/s41598-019-45482-8.
References
- 1.Liverani M, et al. Understanding and managing zoonotic risk in the new livestock industries. Environ Health Perspect. 2013;121:873–877. doi: 10.1289/ehp.1206001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Tang J, et al. Streptococcal Toxic Shock Syndrome Caused by Streptococcus suis Serotype 2. Plos Medicine. 2006;3:e151. doi: 10.1371/journal.pmed.0030151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hričínová M, Holoda E, Mudroňová D, Ondrašovičová S. Multiplex PCR assay for detection of Actinobacillus pleuropneumoniae, Pasteurella multocida and Haemophilus parasuis in lungs of pigs from a slaughterhouse. Folia microbiologica. 2010;55:635–640. doi: 10.1007/s12223-010-0103-9. [DOI] [PubMed] [Google Scholar]
- 4.Kaper JB, Nataro JP, Mobley HLT. Pathogenic Escherichia coli. Nature Review. Nature Reviews Microbiology. 2004;2:123–140. doi: 10.1038/nrmicro818. [DOI] [PubMed] [Google Scholar]
- 5.Mikkelsen LL, Naughton PJ, Hedemann MS, Jensen BB. Effects of Physical Properties of Feed on Microbial Ecology and Survival of Salmonella enterica Serovar Typhimurium in the Pig Gastrointestinal Tract. Applied & Environmental Microbiology. 2004;70:3485. doi: 10.1128/AEM.70.6.3485-3492.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Horiguchi Y. Swine Atrophic Rhinitis Caused by Pasteurella multocida Toxin and Bordetella Dermonecrotic Toxin. Curr Top Microbiol Immunol. 2012;361:113–129. doi: 10.1007/82_2012_206. [DOI] [PubMed] [Google Scholar]
- 7.Brooke CJ, Riley TV. Erysipelothrix rhusiopathiae: bacteriology, epidemiology and clinical manifestations of an occupational pathogen. Journal of Medical Microbiology. 1999;48:789–799. doi: 10.1099/00222615-48-9-789. [DOI] [PubMed] [Google Scholar]
- 8.Gottschalk M, Lacouture S. Canada: Distribution of Streptococcus suis (from 2012 to 2014) and Actinobacillus pleuropneumoniae (from 2011 to 2014) serotypes isolated from diseased pigs. Canadian Veterinary Journal-revue Veterinaire Canadienne. 2015;56:1093–1094. [PMC free article] [PubMed] [Google Scholar]
- 9.Hu X, Zhou Q, Luo Y. Occurrence and source analysis of typical veterinary antibiotics in manure, soil, vegetables and groundwater from organic vegetable bases, northern China. Environmental Pollution. 2010;158:2992–2998. doi: 10.1016/j.envpol.2010.05.023. [DOI] [PubMed] [Google Scholar]
- 10.Ling Z, Hua DY, Hui W. Residues of veterinary antibiotics in manures from feedlot livestock in eight provinces of China. Science of the Total Environment. 2010;408:1069–1075. doi: 10.1016/j.scitotenv.2009.11.014. [DOI] [PubMed] [Google Scholar]
- 11.Barton MD. Impact of antibiotic use in the swine industry. Current Opinion in Microbiology. 2014;19:9–15. doi: 10.1016/j.mib.2014.05.017. [DOI] [PubMed] [Google Scholar]
- 12.Phillips I, et al. Does the use of antibiotics in food animals pose a risk to human health? A critical review of published data. J Antimicrob Chemother. 2004;53:28–52. doi: 10.1093/jac/dkg483. [DOI] [PubMed] [Google Scholar]
- 13.Lun ZR, Wang QP, Chen XG, Li AX, Zhu XQ. Streptococcus suis: an emerging zoonotic pathogen. Lancet Infectious Diseases. 2007;7:201–209. doi: 10.1016/S1473-3099(07)70001-4. [DOI] [PubMed] [Google Scholar]
- 14.Cai X, et al. Serological characterization of Haemophilus parasuis isolates from China. Veterinary microbiology. 2005;111:231–236. doi: 10.1016/j.vetmic.2005.07.007. [DOI] [PubMed] [Google Scholar]
- 15.Tang XB, et al. Isolation, antimicrobial resistance, and virulence genes of Pasteurella multocida strains from swine in China. Journal of clinical microbiology. 2009;47:951. doi: 10.1128/JCM.02029-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bossé JT, et al. Actinobacillus pleuropneumoniae: pathobiology and pathogenesis of infection. Microbes & Infection. 2002;4:225–235. doi: 10.1016/S1286-4579(01)01534-9. [DOI] [PubMed] [Google Scholar]
- 17.Woolfrey BF, Moody JA. Human infections associated with Bordetella bronchiseptica. Clinical Microbiology Reviews. 1991;4:243–255. doi: 10.1128/CMR.4.3.243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chiu CH, Su LC. Salmonella enterica serotype Choleraesuis: epidemiology, pathogenesis, clinical disease, and treatment. Clinical Microbiology Reviews. 2004;17:311. doi: 10.1128/CMR.17.2.311-322.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wood RL. Swine erysipelas–a review of prevalence and research. Journal of the American Veterinary Medical Association. 1984;184:944–949. [PubMed] [Google Scholar]
- 20.Zhao Z, et al. The occurrence of Bordetella bronchiseptica in pigs with clinical respiratory disease. Veterinary Journal. 2011;188:337. doi: 10.1016/j.tvjl.2010.05.022. [DOI] [PubMed] [Google Scholar]
- 21.Delong EF. Archaea in coastal marine environments. Proceedings of the National Academy of Sciences of the United States of America. 1992;89:5685. doi: 10.1073/pnas.89.12.5685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liu Z, et al. Development of Multiplex PCR Assays for the Identification of the 33 Serotypes of Streptococcus suis. Plos One. 2013;8:e72070. doi: 10.1371/journal.pone.0072070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Howell KJ, et al. Development of a Multiplex PCR Assay for Rapid Molecular Serotyping of Haemophilus parasuis. Journal of clinical microbiology. 2015;53:3812–3821. doi: 10.1128/JCM.01991-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schwalbe, R., Steelemoore, L. & Goodwin, A. C. Antimicrobial susceptibility testing protocols. Crc Press (2007).
- 25.Jorgensen J, Ferraro M. Antimicrobial susceptibility testing: general principles and contemporary practices. Clinical Infectious Diseases. 1998;26:973–980. doi: 10.1086/513938. [DOI] [PubMed] [Google Scholar]
- 26.Opriessnig T, Giménez-Lirola LG, Halbur PG. Polymicrobial respiratory disease in pigs. Anim Health Res Rev. 2011;12:133–148. doi: 10.1017/S1466252311000120. [DOI] [PubMed] [Google Scholar]
- 27.Guest JF, Morris A. Community-acquired pneumonia: the annual cost to the National Health Service in the UK. European Respiratory Journal. 1997;10:1530. doi: 10.1183/09031936.97.10071530. [DOI] [PubMed] [Google Scholar]
- 28.Zar HJ, Ferkol TW. The global burden of respiratory disease-impact on child health. Pediatr Pulmonol. 2014;49:430–434. doi: 10.1002/ppul.23030. [DOI] [PubMed] [Google Scholar]
- 29.Katsumi M, Kataoka Y, Takahashi T, Kikuchi N, Hiramune T. Bacterial isolation from slaughtered pigs associated with endocarditis, especially the isolation of Streptococcus suis. Journal of Veterinary Medical Science. 1997;59:75. doi: 10.1292/jvms.59.75. [DOI] [PubMed] [Google Scholar]
- 30.Macinnes, J. I. et al. Prevalence of Actinobacillus pleuropneumoniae, Actinobacillus suis, Haemophilus parasuis, Pasteurella multocida, and Streptococcus suis in representative Ontario swine herds. 72, 242–248 (2008). [PMC free article] [PubMed]
- 31.Moorkamp L, Nathues H, Spergser J, Tegeler R, Grosse BE. Detection of respiratory pathogens in porcine lung tissue and lavage fluid. Veterinary Journal. 2008;175:273–275. doi: 10.1016/j.tvjl.2007.01.008. [DOI] [PubMed] [Google Scholar]
- 32.Lee KE, Choi HW, Kim HH, Song JY, Yang DK. Prevalence and Characterization of Actinobacillus pleuropneumoniae Isolated from Korean Pigs. Journal of Bacteriology & Virology. 2015;45:397–402. [Google Scholar]
- 33.Cowart RP, Bäckström L, Brim TA. Pasteurella multocida and Bordetella bronchiseptica in atrophic rhinitis and pneumonia in swine. Canadian journal of veterinary research = Revue canadienne de recherche vétérinaire. 1989;53:295–300. [PMC free article] [PubMed] [Google Scholar]
- 34.Navacharoen N. Hearing and vestibular loss in Streptococcus suis infection from swine and traditional raw pork exposure in northern Thailand. Journal of Laryngology & Otology. 2009;123:857–862. doi: 10.1017/S0022215109004939. [DOI] [PubMed] [Google Scholar]
- 35.Hill JE, et al. Biochemical analysis, cpn 60 and 16S rDNA sequence data indicate that Streptococcus suis serotypes 32 and 34, isolated from pigs, are Streptococcus orisratti. Veterinary microbiology. 2005;107:63–69. doi: 10.1016/j.vetmic.2005.01.003. [DOI] [PubMed] [Google Scholar]
- 36.Kielstein P, Rappgabrielson VJ. Designation of 15 serovars of Haemophilus parasuis on the basis of immunodiffusion using heat-stable antigen extracts. Journal of clinical microbiology. 1992;30:862–865. doi: 10.1128/jcm.30.4.862-865.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jiang, X. et al. Live Streptococcus suis type 5 strain XS045 provides cross-protection against infection by strains of types 2 and 9. Vaccine34 (2016). [DOI] [PubMed]
- 38.Bak H, Riising H. Protection of vaccinated pigs against experimental infections with homologous and heterologous Hoemophilus parasuis. Veterinary Record. 2002;151:502–505. doi: 10.1136/vr.151.17.502. [DOI] [PubMed] [Google Scholar]
- 39.Wei ZG, et al. Characterization of Streptococcus suis isolates from the diseased pigs in China between 2003 and 2007. Veterinary microbiology. 2009;137:196–201. doi: 10.1016/j.vetmic.2008.12.015. [DOI] [PubMed] [Google Scholar]
- 40.Wang J, et al. Isolation, Identification and Serotyping of Haemophilus parasuis in Parts of Larg-scale Pig Farms in China 2012. Progress in Veterinary Medicine. 2014;35:48–52. [Google Scholar]
- 41.Tadjine M, Mittal KR, Bourdon S, Gottschalk M. Development of a New Serological Test for Serotyping Haemophilus parasuis Isolates and Determination of Their Prevalence in North America. Journal of clinical microbiology. 2004;42:839–840. doi: 10.1128/JCM.42.2.839-840.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Rúbies X, et al. Prevalence of Haemophilus parasuis serovars isolated in Spain from 1993 to 1997. Veterinary microbiology. 1999;66:245. doi: 10.1016/S0378-1135(99)00007-3. [DOI] [PubMed] [Google Scholar]
- 43.Aarestrup FM, Oliver DC, Burch DG. Antimicrobial resistance in swine production. Animal Health Research Reviews. 2008;9:135–148. doi: 10.1017/S1466252308001503. [DOI] [PubMed] [Google Scholar]
- 44.Varela NP, et al. Antimicrobial resistance and prudent drug use for Streptococcus suis. Animal health research reviews. 2013;14:68–77. doi: 10.1017/S1466252313000029. [DOI] [PubMed] [Google Scholar]
- 45.Yang, S. S., Sun, J., Fan, K. W., Yang, X. Y. & Liao, X. P. Investigation of antimicrobial resistance and genetic correlation analysis of Haemophilus parasuis isolated from Guangdong Province. Chinese Veterinary Science (2016).
- 46.Hu P, et al. Comparative genomics study of multi-drug-resistance mechanisms in the antibiotic-resistant Streptococcus suis R61 strain. PLoS One. 2011;6:e24988. doi: 10.1371/journal.pone.0024988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Okwumabua O, O’Connor M, Shull E. A polymerase chain reaction (PCR) assay specific for Streptococcus suis based on the gene encoding the glutamate dehydrogenase. Fems Microbiology Letters. 2003;218:79–84. doi: 10.1111/j.1574-6968.2003.tb11501.x. [DOI] [PubMed] [Google Scholar]
- 48.Oliveira S, Galina L, Pijoan C. Development of a PCR Test to Diagnose Haemophilus Parasuis Infections. Journal of Veterinary Diagnostic Investigation. 2001;13:495–501. doi: 10.1177/104063870101300607. [DOI] [PubMed] [Google Scholar]
- 49.Nagai S, Someno S, Yagihashi T. Differentiation of toxigenic from nontoxigenic isolates of Pasteurella multocida by PCR. Journal of clinical microbiology. 1994;32:1004–1010. doi: 10.1128/jcm.32.4.1004-1010.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bej AK, Mccarty SC, Atlas RM. Detection of coliform bacteria and Escherichia coli by multiplex polymerase chain reaction: comparison with defined substrate and plating methods for water quality monitoring. Applied & Environmental Microbiology. 1991;57:2429. doi: 10.1128/aem.57.8.2429-2432.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gram T, Ahrens P, Andreasen M, Nielsen JP. An Actinobacillus pleuropneumoniae PCR typing system based on the apx and omlA genes–evaluation of isolates from lungs and tonsils of pigs. Veterinary microbiology. 2000;75:43–57. doi: 10.1016/S0378-1135(00)00206-6. [DOI] [PubMed] [Google Scholar]
- 52.Hozbor D, Fouque F, Guiso N. Detection of Bordetella bronchiseptica by the polymerase chain reaction. Research in Microbiology. 1999;150:333–341. doi: 10.1016/S0923-2508(99)80059-X. [DOI] [PubMed] [Google Scholar]
- 53.Chiu CH, Ou JT. Rapid identification of Salmonella serovars in feces by specific detection of virulence genes, invA and spvC, by an enrichment broth culture-multiplex PCR combination assay. Journal of clinical microbiology. 1996;34:2619–2622. doi: 10.1128/jcm.34.10.2619-2622.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Shimoji Y, Mori Y, Hyakutake K, Sekizaki T, Yokomizo Y. Use of an Enrichment Broth Cultivation-PCR Combination Assay for Rapid Diagnosis of Swine Erysipelas. Journal of clinical microbiology. 1998;36:86. doi: 10.1128/jcm.36.1.86-89.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.




