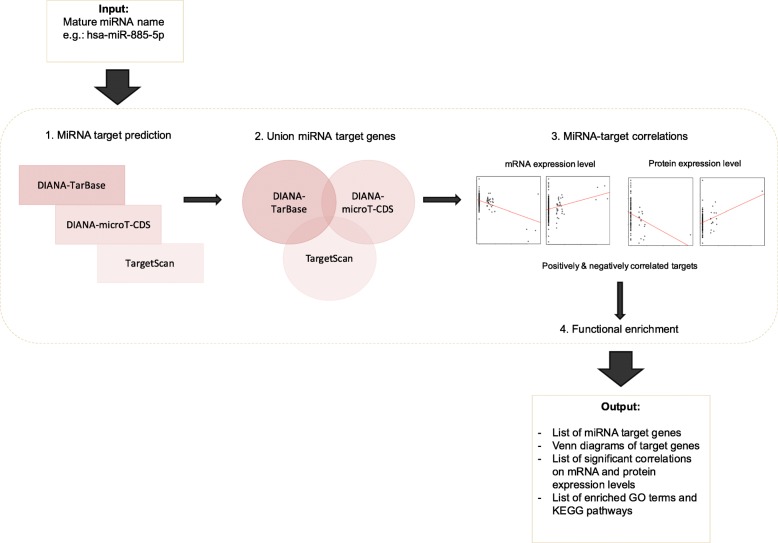
Fig. 1.
Overview of miRFA pipeline. The input is a mature miRNA name. MiRNA target prediction is performed in DIANA-Tarbase v7, DIANA-microT-CDS and TargetScan v7.1 (1.). The union of predicted miRNA targets (2.) were established as well as correlation values for miRNA-mRNA and miRNA-protein expression (3.). The list of correlated miRNA targets was subjected to functional enrichment analysis (4.) for gene ontology (GO) terms and Kyoto encyclopedia of genes and genomes (KEGG) pathways. The output is a list of miRNA target genes, Venn diagrams of target genes, significantly correlated target genes on mRNA and protein expression levels, and enriched GO terms and KEGG pathways