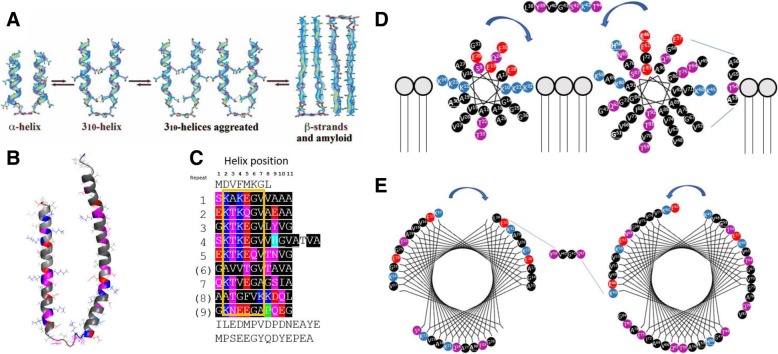
Fig. 4.
The KTKEGV imperfect repeats found within the αS structure. a Model of conformational transition proposed by Singh et al. [47] (CC BY-NC 4.0) of the transition of a 4-peptide bundle into amyloid fibrils, from an alpha-helix into a β-sheet fibril via aggregation induced stabilisation of anti-parallel 310-helix bundles. This model may be representative of the transitions which occur with aS from an alpha–helix membrane bound monomer to β-sheet fibril. b Structure of the micelle bound human aS, published by Ulmer et al., determined by solution NMR spectroscopy [99], highlighting the antiparallel α-helices of the membrane bound αS monomer, helix 1 spanning from Val [3]-Val [38] and helix 2 spanning from Lys [46]-Thr [93], connected by a well ordered linker. c The linear 140 residues of human aS arranged into KTKEGV imperfect repeats 1–9. Blue = basic; light blue = his; red = acidic; purple = polar uncharged; black = nonpolar. d Shown is a colour coded schematic with repeats 1–7 arranged into two 11/3 helix (3 turns over 11 residues), adapted from the αS helical wheels proposed by Dettmar 2018 [100] and Bendor et al. 2013 [101] representative of the membrane induced amphipathic helix. It has been proposed that lysine rich positions (blue) interact with negatively charged lipid head groups, while hydrophobic regions (black, grey area) interact with membrane lipids. Interestingly the Gly residues are found at the hydrophobic-water boundaries of the core, and are found on the adjacent helix face, which may be important in facilitating alpha to β switching at the water membrane, as previously seen in amyloid beta [102]. The position of single amino acid changes associated with early onset PD mutations might destabilise sidechain-sidechain packing that promotes formation of the helix and thereby accelerate the pathway toward amyloidosis. e Proposed structure of 2 × 310 helical wheel, formed by constriction of the α–helical domains seen in the micelle structure, clearly shows that the separation of the Lys and Glu residues in the aS amino acid sequence causes then to stack on top of each other stabilising the 310 intermediate, driving the energetic landscape towards the β-sheet fibril. Most interesting here is that the the first of the ‘ionic locks’ observed in the cryoEM structures is already formed in this structure, between K58-E61. In this proposed structure there does not appear to be a membrane binding domain. Potentially this structural change from α-helix to 310 intermediate could cause membrane disruption and mediate toxicity of αS