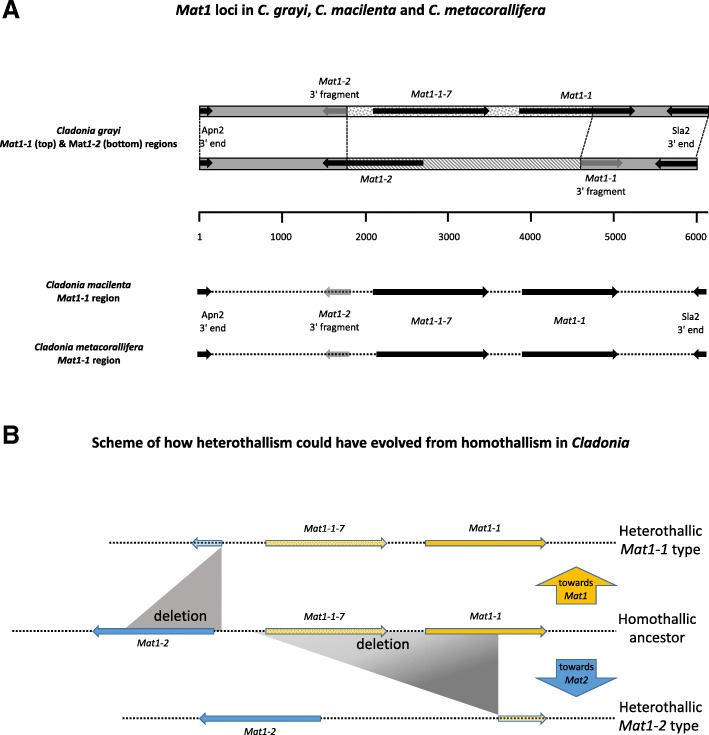
Fig. 4.
Cladonia MAT loci and their evolution. a Configurations of the MAT loci in three Cladonia species. The top diagrammed alignment is based on the alignment between the annotated C. grayi MAT1–1 region (scaffold_00075:76000–90,000 at [67]) and a provisional sequence of the C. grayi MAT1–2 region (accession MH795990). The C. grayi MAT1–1 and MAT1–2 regions are drawn above the basepair indicator line. Under the line are the MAT1–1 regions derived from the genomes of two other Cladonia species (Additional file 5). In C. grayi, the conserved flanking regions are gray, while the unrelated central regions are stippled differently for each mating type. Dark or gray arrows represent genes and gene-segments. CLAGR_008123-RA is considered a putative MAT1–1-7 ortholog because of its location and its BLAST hits to MAT1–1-7 orthologs from Trichophyton and other fungi. b Evolutionary model. Horizontal colored arrows represent MAT idiomorphs. The central line represents the MAT locus configuration of a possible homothallic Cladonia ancestor, and the vertical arrows represent the putative transitions towards the present heterothallic MAT1–1 (orange) or MAT1–2 (blue) configurations (Additional file 5). The graded shading in the deletion triangle leading to MAT1–2 symbolizes the deletion’s undefined left boundary beyond MAT1–1-7