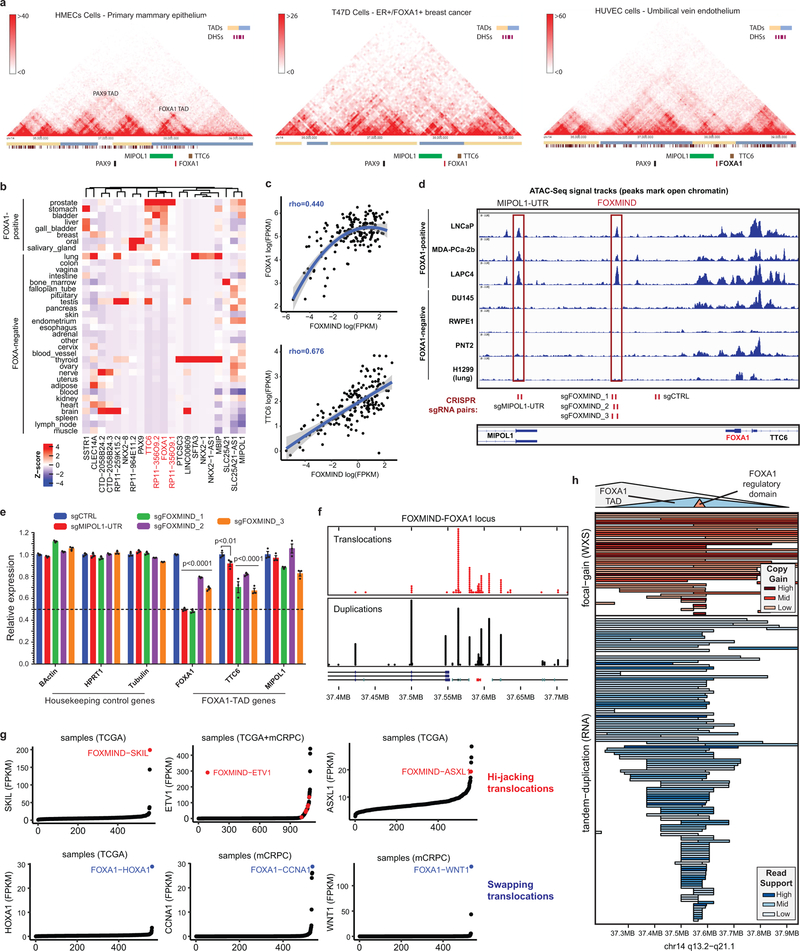
Extended Data Figure 9. Topological, physical, and transcriptional characteristics of the FOXA1 locus in normal tissues and prostate cancer.
a) HI-C data (from: http://promoter.bx.psu.edu/hi-c/view.php) depicting conserved topological domains within the PAX9/FOXA1 syntenic block in normal and FOXA1-positive cancer cell lines. b) Highly tissue-specific patterns of gene expression within the PAX9/FOXA1 syntenic block. Tissues were dichotomized into FOXA1+ and FOXA1- based on FOXA1 expression levels; genes were subject to unsupervised clustering. Z-score normalization was performed for each gene across all tissues. c) Correlation of FOXMIND (Methods) and FOXA1 / TTC6 expression levels across metastatic tissues (n=370; Spearman rank-correlation coefficient). The 95% confidence interval is shown. d) Representative ATAC-seq (n=1) read signal tracks from normal basal epithelial prostate (RWPE1, PNT2) or PCa cells. Cells are grouped based on expression of FOXA1 and differentially pioneered loci are marked with the red boxes. CRISPR sgRNA pairs used for genomic deletion of the labelled elements are shown at the bottom. Distinct FOXA1+ and FOXA1- cell lines serve as biological replicates for ATAC-seq. e) mRNA (qPCR) expression of control, FOXA1 TAD genes, and MIPOL1 in VCaP cells treated with CRISPR-sgRNA pairs targeting a control site (sgCTRL), the FOXMIND, or the MIPOL1-UTR regulatory element (see Extended Data Fig. 2c for sgRNA binding sites). Distinct sgRNA pairs cutting at FOXMIND serve as biological replicates. Mean ± s.e.m are shown (n=3 technical replicates; two-way ANOVA and Tukey’s test). f) Distribution of tandem duplication and translocation breakends (chimeric junctions or copy-number segment boundaries) focused at the FOXMIND-FOXA1 regulatory domain. g) Outlier expression of genes involved in translocations with the FOXA1 locus. Translocations positioning a gene between FOXMIND and FOXA1 (Hi-jacking) are shown on top (red). Translocations positioning a gene upstream of the FOXA1 promoter (Swapping) are shown on the bottom (blue). h) Inferred duplications within the FOXA1 locus based on RNA-seq (tandem breakends) and WES (copy-gains) zoomed-in at the FOXA1 TAD.