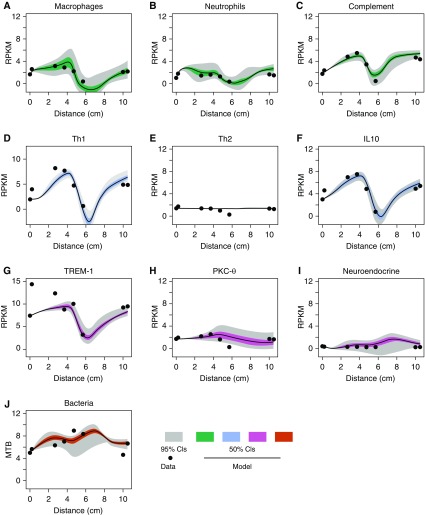
Figure 5.
Model fit for dynamical sink with coupled interaction network. Model fits are shown with mean per kilobase per million mapped read (RPKM) values for each module (consisting of all genes in the module), for (A) macrophages, (B) neutrophils, (C) complement, (D) Th1, (E) Th2, (F) IL-10, (G) TREM-1, (H) protein kinase C-θ, (I) neuroendocrine, and (J) Mycobacterium tuberculosis burden. There were deep wells for the sink encountered for macrophages (A), complement (C), Th1 (D), IL-10 (F), and TREM-1 (G) at 6–8 cm. The neuroendocrine system/pathway looks flat because of the scale used and the relatively low RPKM reads (mostly downregulated compared with nontubersulosis lung). The scaled-up figure is shown in Figure E8. Table E3 shows the dynamical sink parameters of the neuroendocrine module. CI = confidence interval; Mtb = Mycobacterium tuberculosis; PKCθ = protein kinase C-θ; Th1 = T-helper cell type 1; TREM-1 = triggering receptor expressed on myeloid cells-1.