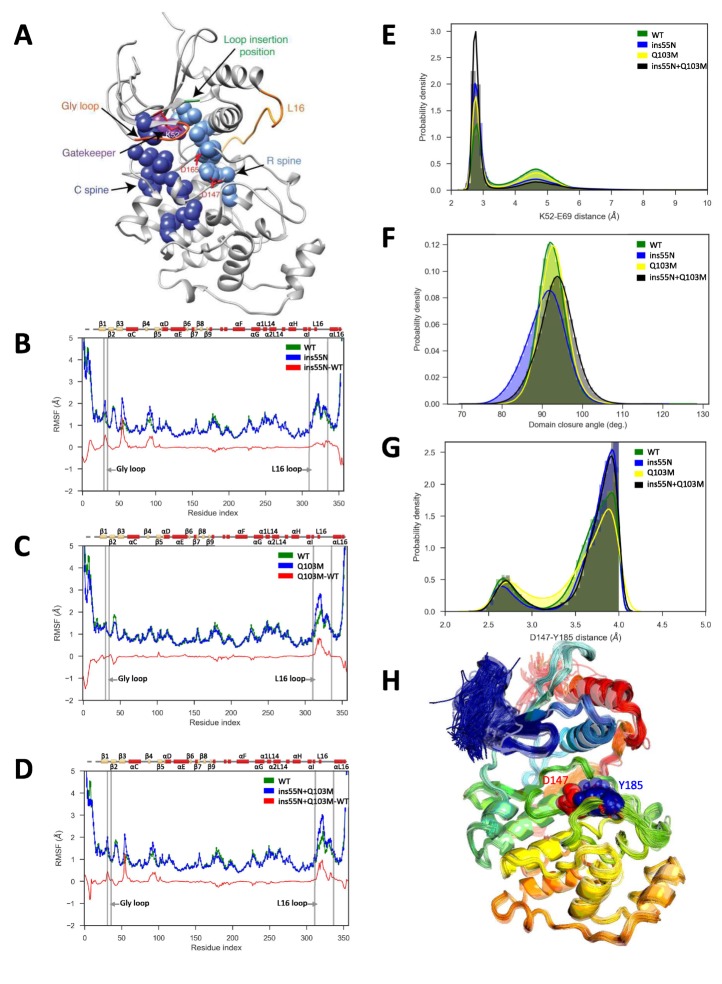
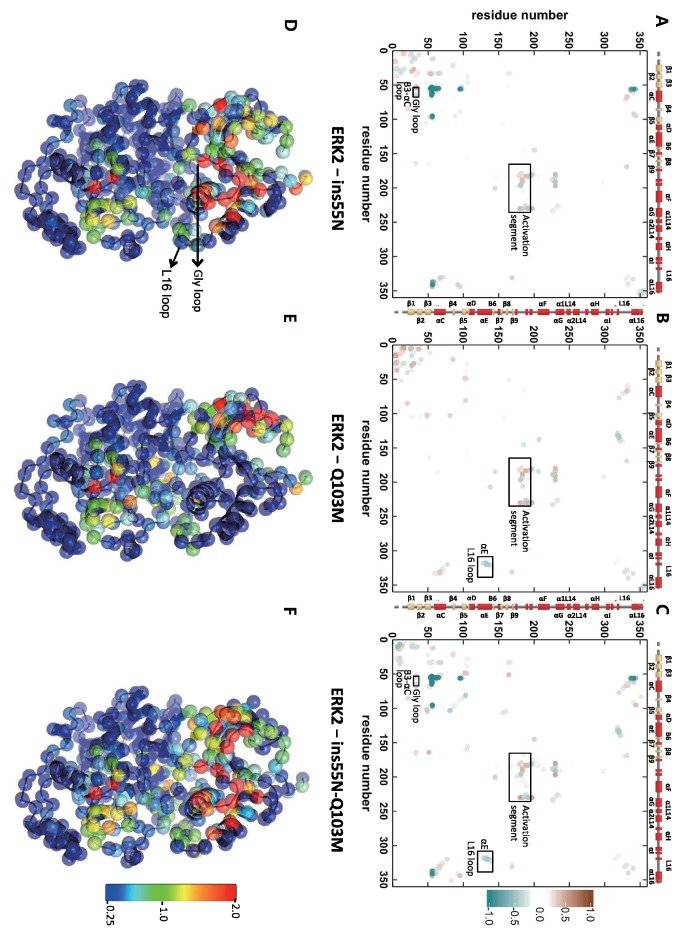
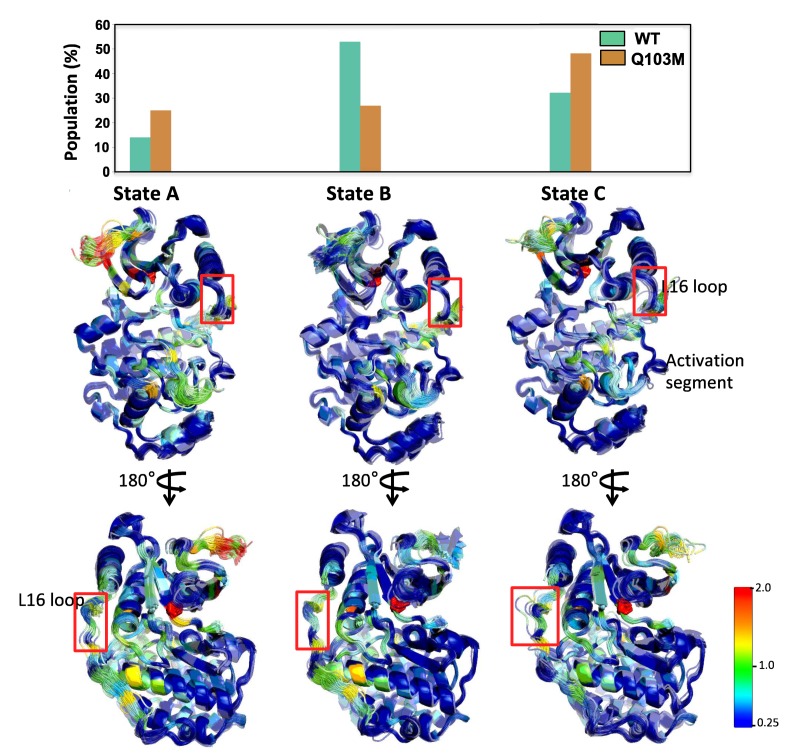
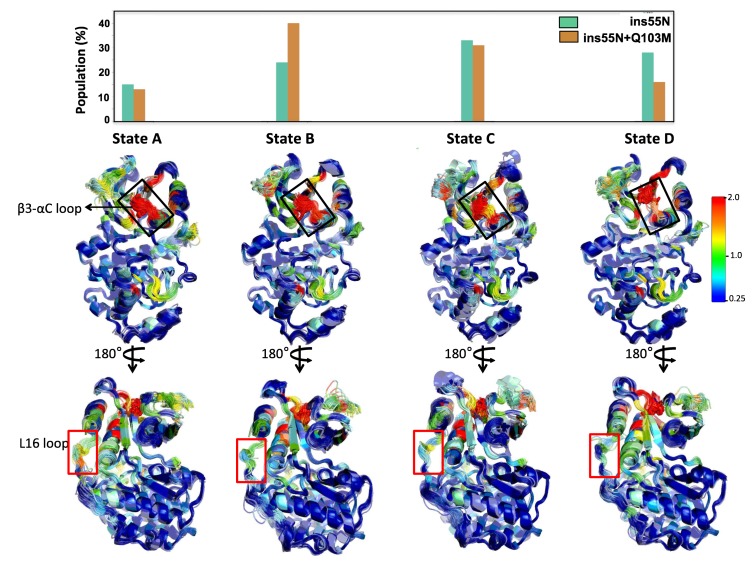
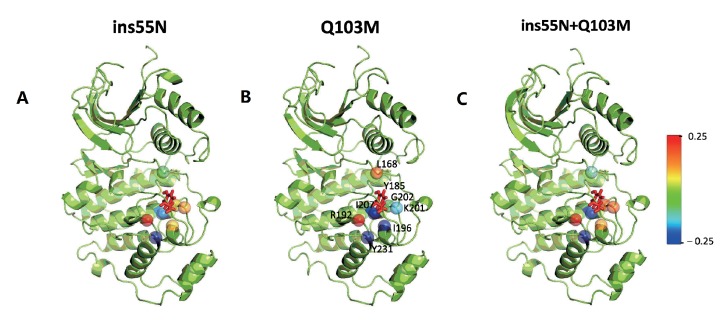
Figure 5. Molecular dynamics simulations of activated mutants show increased flexibility at the glycine rich loop and loop 16, and increased salt-bridge formation at the catalytic site.
(A) Structure of ERK2 with relevant features annotated: L16 – orange, Gly loop - dark orange, C spine - dark blue, R spine - light blue, gatekeeper residue – purple, catalytic residues (K52, D147 and D165) – red, and loop insertion position – green. (B–D) Root mean square fluctuations (RMSF, error bars indicating 95% confidence intervals of the means are plotted but are too small to see): Wild type – green, mutant – blue and difference between mutant and WT – red. (B) ERK2-ins55N (ß3-αC loop insertion mutant) (C) ERK2-Q103M (gatekeeper residue mutant) (D) ERK2-ins55N-Q103M (double mutant). (E) Probability density of salt-bridge between ß3-lysine and αC-glutamate, K52–E69. (F) Distribution of domain closure angles. (G) Probability density of distances < 4 Å between Y185 of the ‘TxY’ motif and active site D147H: Wild type – green, ERK2-ins55N – blue, ERK2-Q103M – yellow and ERK2-ins55N-Q103M – black. (H). Top 100 conformations of ERK2-WT with smallest Y185-D147 distance. D147 (red) and Y185 (blue) are shown as spheres.