Abstract
Background
Interpretative reading of antimicrobial susceptibility test (AST) results allows inferring biochemical resistance mechanisms from resistance phenotypes. For aminoglycosides, however, correlations between resistance pathways inferred on the basis of the European Committee on Antimicrobial Susceptibility Testing (EUCAST) clinical breakpoints and expert rules versus genotypes are generally poor. This study aimed at developing and validating a decision tree based on resistance phenotypes determined by disc diffusion and based on epidemiological cut-offs (ECOFFs) to infer the corresponding resistance mechanisms in Escherichia coli.
Methods
Phenotypic antibiotic susceptibility of thirty wild-type and 458 aminoglycoside-resistant E. coli clinical isolates was determined by disc diffusion and the genomes were sequenced. Based on well-defined cut-offs, we developed a phenotype-based algorithm (Aminoglycoside Resistance Mechanism Inference Algorithm - ARMIA) to infer the biochemical mechanisms responsible for the corresponding aminoglycoside resistance phenotypes. The mechanisms inferred from susceptibility to kanamycin, tobramycin and gentamicin were analysed using ARMIA- or EUCAST-based AST interpretation and validated by whole genome sequencing (WGS) of the host bacteria.
Findings
ARMIA-based inference of resistance mechanisms and WGS data were congruent in 441/458 isolates (96·3%). In contrast, there was a poor correlation between resistance mechanisms inferred using EUCAST CBPs/expert rules and WGS data (418/488, 85·6%). Based on the assumption that resistance mechanisms can result in therapeutic failure, EUCAST produced 63 (12·9%) very major errors (vME), compared to only 2 (0·4%) vME with ARMIA. When used for detection and identification of resistance mechanisms, ARMIA resolved >95% vMEs generated by EUCAST-based AST interpretation.
Interpretation
This study demonstrates that ECOFF-based analysis of AST data of only four aminoglycosides provides accurate information on the resistance mechanisms in E. coli. Since aminoglycoside resistance mechanisms, despite having in certain cases a minimal effect on the minimal inhibitory concentration, may compromise the bactericidal activity of aminoglycosides, prompt detection of resistance mechanisms is crucial for therapy. Using ARMIA as an interpretative rule set for editing AST results allows for better predictions of in vivo activity of this drug class.
Research in context.
Evidence before study
Interpretative reading of antibiogram data allows to infer underlying resistance mechanisms and thus better prediction of in vivo efficacy of drugs. EUCAST guidelines include a number of expert rules, screening cut-off values and diagnostic algorithms to infer resistance mechanisms and to review AST results. However, while these methods allow reliable and accurate detection of resistance pathways for some drug classes, such as β-lactams, for others, such as aminoglycosides, phenotypic analyses often result in poor genotype/phenotype correlations. This is due, at least in part, to the fact that the interpretation is mostly based on clinical breakpoints (CBPs), which take into account parameters unrelated to in vitro susceptibility testing. In contrast to CBPs, epidemiological cut-offs (ECOFFs) can precisely and in an unbiased manner separate the wild-type microorganisms without acquired resistance mechanisms from non-wild-type isolates. The use of ECOFFs in the interpretation of AST data can conceptually help to resolve the inherent limitations of the CBPs-based approach.
Added value of this study
Using both epidemiological cut-offs (ECOFFs) and WGS-based genotypic data, we built a phenotype based diagnostic algorithm (Aminoglycoside Resistance Mechanism Inference Algorithm - ARMIA) allowing reliable detection of aminoglycoside resistance mechanisms in Escherichia coli. Agreement between the mechanisms inferred by ARMIA and WGS was of 96·5%. Similarly, the concordance between WGS-based genotypic data and the expected phenotypes was of 97·5%. In contrast, congruence between resistance mechanisms inferred on the basis of EUCAST interpretation criteria and genotypic data was of only 85·7%. Based on the assumption that resistance genes can result in clinical resistance, EUCAST- and ARMIA-based analyses produced 63 (12·9%) and 2 (0·4%) very major errors (vME), respectively. When used as an expert rule to infer resistance mechanisms, ARMIA resolved 95% of the vMEs generated by EUCAST-based AST interpretation.
Implication of all available evidence
In this study we demonstrate how a detailed analysis of phenotype-based antimicrobial susceptibility testing data of only four aminoglycosides can provide reliable information on resistance mechanisms. Since aminoglycoside-modifying enzymes, despite causing in certain cases a slight effect on the aminoglycoside minimal inhibitory concentrations, can significantly affect their bactericidal activity, their detection is crucial for the selection of the most appropriate antibiotics. The phenotype-based ARMIA can be used as an expert system to edit AST results for therapeutic predictions as well as for epidemiological studies. ARMIA can be considered as a paradigm for the analysis of other complex antibiotic class resistance patterns.
Alt-text: Unlabelled Box
1. Introduction
Bacterial pathogens are clinically categorized as susceptible or resistant on the basis of antimicrobial susceptibility testing (AST) results. These data can also be used to infer the underlying resistance mechanism(s) by ‘interpretative reading’ of susceptibility patterns [1]. Following this concept, EUCAST has set ‘expert rules’ whereby AST results for one or more ‘indicator drugs’ are used to infer the resistance mechanisms. The ‘indicator drugs’ are usually the best substrates of a given resistance pathway and thus the best suited to detect in vitro resistance. For some antibiotic classes, such as β-lactams, EUCAST provides screening cut-offs and/or diagnostic algorithms for detection and/or confirmation of resistance mechanisms [2]. In contrast, for other drug classes, such as aminoglycosides, the inference of resistance mechanisms from phenotypes is poorly accurate. This in part relates to the habit that EUCAST expert rules are based on clinical breakpoints (CBPs), which are defined to predict the likelihood for therapeutic success, and therefore take also into account parameters unrelated to antimicrobial resistance mechanisms (e.g. PK/PD parameters, clinical outcome data, etc.). In contrast, epidemiological cut-offs (ECOFFs) are selected to precisely discriminate the wild-type microorganisms without acquired resistance mechanisms from non-wild-type isolates [3].
Precision medicine is a challenge for patient management in infectious diseases. Due to the rapidly falling costs and shortening of the turnaround time, whole genome sequencing (WGS) is increasingly being viewed as a means to improve patient care through detection of mutations or acquired resistance genes [4]. However, despite the interest of the scientific community, implementation in the clinic laboratory has been slow. The major drawback of using WGS-inferred antimicrobial susceptibility testing to guide clinical decision making is the lack of sufficient evidence linking phenotypic and genotypic data [5]. This gap is antibiotic- and/or species-dependent and in general results from combined or not yet reported resistance mechanisms. Despite these limitations, WGS can be used for detection of well described resistance mechanisms. A prominent example is Mycobacterium tuberculosis, which has evolved as a prime target for genotypic testing [[6], [7], [8]].
We investigated the potentials and limitations of phenotype- and genotype-based inference of antibiotic resistance mechanisms. We selected aminoglycoside resistance in Escherichia coli as paradigmatic example for the following reasons: (i) aminoglycosides are ‘must medications’ for empirical combination therapy of serious infections with Gram-negative bacteria [9], (ii) the presence of a large number of extensively studied resistance mechanisms, which are mainly related to the production of aminoglycoside-modifying enzymes (AMEs) and 16S rRNA methyltransferases (RMTases) [10], (iii) the availability of EUCAST ECOFFs and CBPs for most clinically-relevant aminoglycosides [3] and (iv) several studies have reported a surprisingly poor phenotype/genotype correlation, indicating conceptual errors in AST interpretation [[11], [12], [13]].
To date >50 types of AMEs have been described [14]. They are grouped in three classes according to their modifying activities, namely acetyltransferases (AAC), phosphotransferases (APH) and nucleotidyltransferases (ANT). Within these classes the enzymes can modify aminoglycosides at different sites of the drug scaffold. The activity of AMEs is often not restricted to only one aminoglycoside and a given aminoglycoside can be modified by several AMEs. In addition, more than one enzyme can be present in an isolate. Since only a limited number of aminoglycosides is usually tested, inferring AMEs from phenotypic analysis of resistance can be problematic and molecular techniques are often required to identify the underlying mechanisms responsible for the phenotypes [11,15]. To solve these issues Van de Klundert et al. developed in 1984 a highly accurate determination method for the identification of AMEs in clinical isolates based on inhibition zone diameters for six commercially available aminoglycosides [16].
Here, we present ARMIA (Aminoglycoside Resistance Mechanism Inference Algorithm), a dichotomic decision tree for the inference of resistance mechanisms from inhibition zone diameters. The algorithm uses ECOFFs for gentamicin, tobramycin and kanamycin as well as a working separator cut-off for amikacin and was established based on the analysis of the resistance phenotypes and genotypes of 488 E. coli clinical isolates. We compared the performance of ARMIA with that of WGS in predicting aminoglycoside resistance and investigated the potential of this algorithm-based interpretation to resolve EUCAST-based categorization errors.
2. Materials and methods
2.1. Study design
We conducted a laboratory-based, single-centre comparison study of aminoglycoside resistance in E. coli. For reporting, STARD 2015 guidelines were followed [17].
2.2. Clinical isolates
Strains used in this study were selected among the 5575 E. coli clinical isolates collected between February 1, 2014 and December 31, 2014, in the diagnostic laboratory at the Institute of Medical Microbiology (IMM), University of Zurich. Strains were considered duplicates and discarded if they (i) originated from the same patient and (ii) did not show at least one major and two minor differences in AST interpretation (Table S1). Based on disc diffusion data for gentamicin, tobramycin and kanamycin, and epidemiologic cut-off values (EUCAST ECOFFs of 16 mm for tobramycin and gentamicin [18] and the local IMM ECOFF of 15 mm for kanamycin), 3068/3359 non-duplicate E. coli clinical strains were categorized as susceptible to the aminoglycosides tested. All 451 E. coli clinical isolates classified as resistant to at least one aminoglycoside and thirty of the 3068 susceptible strains were included in the study and subjected to WGS. In addition, seven E. coli highly resistant to all aminoglycosides (inhibition zone diameters = 6 mm, collected at the IMM between January 1, 2015 and December 31, 2017), putatively producing RMTases, were added [19].
2.3. Antimicrobial susceptibility testing
Disc diffusion was performed in triplicate according to the EUCAST guidelines [20]. Kanamycin [a mixture of kanamycin B (98%) and kanamycin A (2%)], tobramycin, amikacin, and gentamicin were purchased from Oxoid Limited, Basingstoke, United Kingdom. The Sirweb/Sirscan system (i2a) was used to measure and electronically archive the inhibition zone diameters [21]. To reduce the impact of outliers, the median of the triplicates was considered for the assignment to a resistance phenotype.
2.4. Time-kill assay
Amikacin and kanamycin minimal inhibitory concentrations (MICs) against selected clinical isolates were determined by broth microdilution [22]. Standard inocula of 1 × 106 CFU/ml were challenged with amikacin and kanamycin at 0.5×, 1×, 2×, and 4× MIC levels. Appropriate dilutions were plated and the surviving bacteria were enumerated at time 0 and after 2, 4, 8 and 24 h. Bactericidal activity was defined as ≥3 log decrease in CFU/ml after 24 h of incubation.
2.5. Whole genome sequencing
Library preparation for whole genome sequencing was performed with the QIAGEN QIASeq FX kit. The quality of the library was assessed using capillary electrophoresis (Fragment Analyzer Automated CE System, Advanced Analytical, Heidelberg, Germany). DNA libraries were pooled in equal concentrations and paired-end sequencing (2 × 150) was performed on an Illumina MiSeq platform (Illumina, San Diego, CA, USA). Sequences were deposited in the European Nucleotide Archive under accession number PRJEB29576.
2.6. Detection of resistance genes
Raw sequencing reads (FASTQ) were filtered and trimmed using the fastq trimmer tool of the FASTX-Toolkit (Hannon Lab, Cold Spring Harbour Laboratories, New York, NY, USA) applying a threshold PHRED score ≤ 25. To identify antibiotic resistance genes and mutations, trimmed FASTQ reads were analysed using the ARIBA pipeline [23], querying the ARG-ANNOT [24] and CARD databases [25].
2.7. Multi locus sequence typing (MLST)
MLST according to the “Warwick” scheme [26] was performed using the Ridom Seqsphere+ software (Ridom Bioinformatics, Münster, Germany).
2.8. Statistical method
All calculations were performed with the software R, version 3.2.3 (freely available under http://www.r-project.org/) [27]. Sensitivity of ARMIA was determined by dividing the sum of strains with concordant phenotype/genotype and discrepants that are phenotypically resistant and genotypically susceptible (likely resulting from deficiencies of the genetic testing, for explanation see the Result section) by the total number of strains. Specificity of ARMIA was calculated by dividing the number of strains with concordant genotype/phenotype by the total number of tested clinical isolates.
2.9. Ethics
The research was conducted in accordance with the Declaration of Helsinki and national and institutional standards. Ethical approval was not required since only anonymized health related data (i.e. antibiotic susceptibility results) of clinical isolates were used. The Swiss Human research Act does not apply to this study.
3. Results
The aminoglycoside susceptibility profiles of thirty wild-type and 458 E. coli clinical strains resistant to at least one among gentamicin, tobramycin or kanamycin were studied [20]. Based on information derived from the literature (Table S2), ECOFFs for gentamicin, tobramycin and kanamycin, and a working separator cut-off for amikacin, we developed a phenotype-based algorithm (Aminoglycoside Resistance Mechanism Inference Algorithm, hereafter ARMIA) to infer aminoglycoside resistance mechanisms and thus predict clinical resistance to aminoglycosides (Fig. 1). Although it is not used in clinic, kanamycin was selected since it is the best substrate and therefore the most adequate to test the in vitro activity of APH(3′) and AAC(6′) enzymes. The design of the algorithm greatly benefited from the combined analysis of inhibition diameters and genome sequences of the strains. For instance, diameter distributions associated with the various resistance mechanisms (Fig. 2) showed that gentamicin was the best drug to separate the wild-type and non-wild-type populations and it was therefore set as the first node in the flowchart. Moreover, using a working separator cut-off of 22 mm, rather than the EUCAST ECOFF of 18 mm, isolates with AME combinations affecting amikacin could be efficiently separated from those with combinations not affecting amikacin. We found a high agreement (96·3%) between the resistance mechanisms inferred from the observed phenotypes and the WGS-based genotypes. To define the possible clinical consequences resulting from disagreement, i.e. minor errors (mE), major errors (ME) or very major errors (vME), we followed the rule that resistance (phenotypically observed or predicted from WGS) overtakes susceptibility. This was based on two assumptions. First, in in vitro susceptible strains with a genotype predicting resistance, the disagreement likely reflects the failure of phenotypic testing to detect resistance, e.g. due to low-level expression of the corresponding gene [28]. Consequently, these discrepancies are categorized as phenotypic errors. Second, in in vitro resistant isolates with a WGS-based predicted susceptible phenotype, the disagreement was considered to reflect an unknown/yet undescribed resistance mechanism, the strains should thus be considered as resistant and the corresponding discrepancies are categorized as errors of the genetic testing. Applying these rules, only two clinical isolates (0·4%) were classified as vMEs (Table S3). Discrepancies were mostly due to diameter values close to the ECOFFs (Fig. S1, Table S4), suggesting that these unexpected phenotypes may result from activation of intrinsic low-level resistance mechanisms (e.g. increased expression of efflux pumps) rather than AMEs [29].
Fig. 1.
Aminoglycoside Resistance Mechanism Inference Algorithm (ARMIA). AME, aminoglycoside-modifying enzyme; AMK, amikacin (30 μg); GEN, gentamicin (10 μg); RMT: 16S rRNA methyltransferase; TOB, tobramycin (10 μg); R, resistant; S, susceptible. Strains were sequentially divided based on the size of the inhibition zone diameters for the aminoglycosides indicated in the nodes. At the bottom of the diagram the observed phenotypes, from which the underlying resistance mechanisms were inferred, are summarized. For each resistotype, the percentage of clinical isolates with congruent ARMIA-based resistance mechanism(s) and WGS-based genotype is indicated. Predicted phenotype and the clinical impact of discrepancies are displayed at the bottom of the algorithm.
1AMK is not affected by APH(3′)-I,-II, -IV, -V.
2The amikacin working separator cut-off of 22 mm was used to distinguish AME combinations without and with AAC(6′)-I enzymes.
3AAC(3) bearing isolates cannot be separated from AME combinations.
4High-level resistance (diameter = 6 mm) to GEN, KAN and AMK.
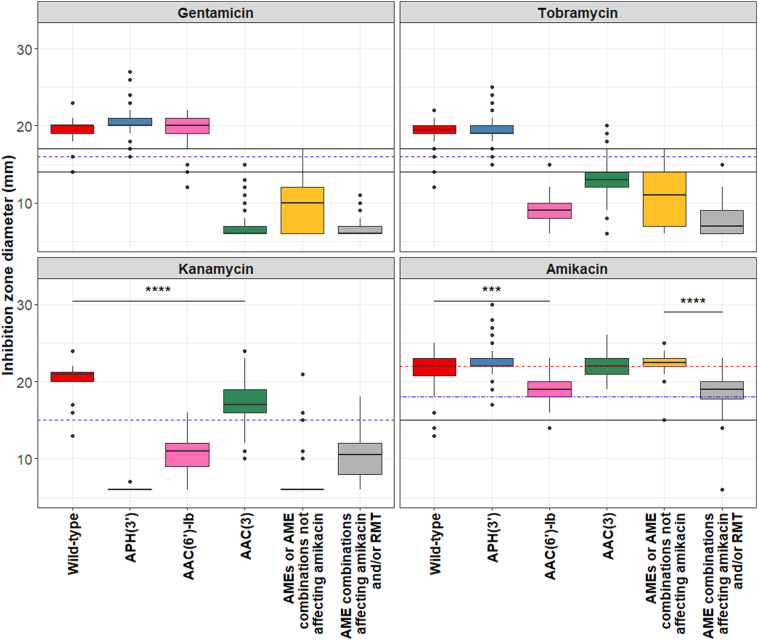
Fig. 2.
Analysis of inhibition zone diameters. Boxplots of growth inhibition diameters. The bottom and top of the box represent the first and third quartiles, whiskers have a maximum of 1.5 IQR, dots represent data not included between the whiskers and bold lines indicate median diameter values. P values <0.001 are summarized with three asterisks and P values <0.0001 are summarized with four asterisks. P values remained highly significant after comparison multiple testings. Solid black lines indicate EUCAST CBPs. Dashed blue lines indicate EUCAST ECOFFs for gentamicin and tobramycin and the IMM ECOFF for kanamycin. The dot dashed blue line indicates superimposition of the EUCAST ECOFF and the upper CBP for amikacin. The red dashed line indicates the working separator cut-off for amikacin. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
We then investigated how WGS-based genotypic data can predict ECOFF-based resistance phenotypes. Based on the genotypes, the strains were grouped by resistance mechanisms or combinations thereof conferring the same resistance phenotypes (referred to as resistotypes) (Fig. 3). If no resistance genes were detected, the isolates were categorized as wild-type. Relating the genetic resistance mechanisms to the corresponding population-based distribution of inhibition zone diameters pointed to two limitations of phenotypic testing. First, for AAC(6′)-producing strains and amikacin, the inhibition zone diameters were close to the ECOFF (Fig. 2). However, population analysis demonstrated that AAC(6′) enzymes affect amikacin, as there is a significant shift in the median diameter from 22 mm (wild-type) to 19 mm [AAC(6′) strains] (Table 1). Second, for AAC(3) strains, the diameters for kanamycin and tobramycin were close to the ECOFFs or CBPs (Fig. 2). Again, population analysis showed that the AAC(3) enzymes affect both kanamycin and tobramycin, as there is a significant reduction in the median diameter for both aminoglycosides (4 mm for kanamycin and 6.5 mm for tobramycin, Table 1). We thus concluded that amikacin testing is not reliable to detect the presence of AAC(6′) enzymes and that tobramycin and kanamycin are not suitable to infer the production of AAC(3). Consequently, we defined the following rules: (i) in the presence of AAC(6′) amikacin testing should be disregarded, (ii) in the presence of AAC(3) tobramycin and kanamycin testing should be disregarded and (iii) the corresponding isolates should be considered resistant to these drugs. These rules were applied in the analysis of WGS predicted phenotypes and ECOFF-based observed phenotypes (Fig. 3, Table S5). Overall, WGS prediction of resistance resulted in vMEs in eight isolates (1·6%) (Table S6). Again, discrepancies were mostly caused by inhibition zone diameters close to the ECOFFs (Fig. S2).
Fig. 3.
WGS-based aminoglycoside resistance phenotypes. AME, aminoglycoside-modifying enzyme; AMK, amikacin (30 μg); GEN, gentamicin (10 μg); RMT: 16S rRNA methyltransferase; TOB, tobramycin (10 μg); R, resistant; S, susceptible. Strains were categorized by resistance mechanism(s) (also referred to as resistotype) on the basis of the WGS data. For each resistotype, the percentage of clinical isolates with congruent predicted and observed phenotype was determined. The clinical impact of the discrepancies is indicated at the bottom.
1Tobramycin and kanamycin testing must be disregarded in the presence of AAC(3) enzymes. 2Amikacin testing must be disregarded in the presence of AAC(6′) enzymes.
Table 1.
Medians (mm) of inhibition zone diameter distributions.
| Aminoglycoside | wt | APH(3′)a | ∆wt/APH(3′) | AAC(3)b | ∆wt/AAC(3) | p-Value | AAC(6′)c | ∆wt/AAC(6′) | p-Value |
|---|---|---|---|---|---|---|---|---|---|
| Gentamicin | 20 | 20 | – | 6 | 14 | 20 | – | ||
| Tobramycin | 19.5 | 19 | 0.5 | 13 | 6.5 | 9 | 10.5 | ||
| Kanamycin | 21 | 6 | 15 | 17 | 4 | <0.0001 | 11 | 10 | |
| Amikacin | 22 | 22 | – | 22 | – | 19 | 3 | 0.0004 |
APH(3′) affects kanamycin but not gentamicin, tobramycin and amikacin.
AAC(3) affects gentamicin, tobramycin, kanamycin but not amikacin.
AAC(6′) affects amikacin, kanamycin, tobramycin but not gentamicin.
Aminoglycoside-resistance mechanisms of the 488 E. coli clinical isolates were then inferred on the basis of EUCAST CBPs and expert rules and confronted to WGS data (Fig. S3, Table S7). Overall, we found a poor correlation (85·9%) between EUCAST AST-interpretation and WGS data. This was in particular the case for strains with AME combinations containing AAC(6′). These results confirm those of previous studies, where high disagreement rates between aminoglycoside resistance mechanisms predicted from phenotypic analyses based on EUCAST or CLSI CBPs and the corresponding genotypes have been observed [[11], [12], [13]]. The potential clinical consequences resulting from disagreement revealed that 63/488 (12·9%) clinical isolates were classified as false susceptible (vME) on the basis of EUCAST CBPs and expert rules (Fig. S3, Table S7).
Given the high reliability of ARMIA to infer biochemical mechanisms, we asked whether ARMIA could resolve EUCAST phenotype/genotype inconsistencies. We thus used ARMIA-based prediction to ‘rule in’ resistance in the clinical isolates classified as susceptible by EUCAST AST interpretation. Overall, ARMIA solved 60/63 vME and one out of three mE in 66 clinical isolates incorrectly categorized following EUCAST guidelines (Table S7). However, ARMIA introduced four ME and three mE in seven strains correctly interpreted by EUCAST. As a consequence, reviewing EUCAST aminoglycoside susceptibility phenotypes by ARMIA interpretative rules would have discouraged the use of these drugs against pathogens with undetected drug resistance genes in 60/63 cases (95·2%).
As shown by population analysis, despite the fact that AAC(6′) and AAC(3) can cause a significant reduction in susceptibility towards amikacin and kanamycin, in most of the cases this does not translate into a change of clinical category (Fig. 2). To assess the impact of AAC(6′) and AAC(3) on the in vitro activity of amikacin and kanamycin against E. coli, time-kill studies were performed. For the experiments with amikacin, two susceptible E. coli strains with no known aminoglycoside resistance genes, one with aac(3)-IId, one with aph(3)-Ia and three strains harbouring the aac(6′)-Ib-cr gene were analysed. Although AAC(6′)-Ib-cr enzymes are known to be less effective against aminoglycosides compared to other members of the same subclass [30], we chose clinical isolates carrying an aac(6′)-Ib-cr gene since this was the most prevalent variant found in the strains tested (109/115, 94·7%). Of note, all E. coli strains had the same amikacin MIC of 8 mg/L and based on EUCAST CBPs (S ≤ 8; R > 16 mg/L) would have been categorized as susceptible. However, while amikacin was bactericidal at 2× MIC against the wild-type strains and the AAC(3)-IId and APH(3)-Ia-producers, no bactericidal activity was observed when exposing AAC(6′)-Ib strains to up to 4× the amikacin MIC (Table 2, Fig. S4). For the kanamycin experiments, two susceptible E. coli strains with no known resistance mechanisms and two AAC(3)-II producers were studied. All the isolates had the same MIC, 16 mg/L, and would thus be classified as susceptible according to the CLSI recommendations (S ≤ 16; R ≥ 64 mg/L). However, while 1× MIC was bactericidal against the two wild-type strains, over 4× the MIC was not bactericidal against the two AAC(3)-II producers (Table 3, Fig. S5).
Table 2.
AAC(6′)-Ib compromises amikacin bactericidal activity.
| No | Resistotype | Amikacin |
Log kill at 24 h (CFU/ml) |
|||
|---|---|---|---|---|---|---|
| MIC | 0.5 x MIC | 1 x MIC | 2 x MIC | 4 x MIC | ||
| 1 | Wild-type | 8 | 2.3 | 2.2 | −4.8 | −6.4 |
| 2 | Wild-type | 8 | 1.8 | 1.1 | −6.2 | −6.3 |
| 3 | AAC(3)-IId | 8 | 1.6 | 1.9 | −6.3 | −6.4 |
| 4 | APH(3′)-Ia | 8 | 2.3 | −0.4 | −6.2 | −6.1 |
| 5 | AAC(6′)-Ib-cr | 8 | 2.5 | 2.6 | 2.6 | 2.3 |
| 6 | AAC(6′)-Ib-cr | 8 | 2.5 | 2.5 | 0.7 | 0.6 |
| 7 | AAC(6′)-Ib-cr | 8 | 2.8 | 2.7 | 2.7 | 2.6 |
Boldface red numbers indicate bactericidal activity (≥ 3 log decrease of the bacterial load from the initial inocolum after 24 h of incubation).
Table 3.
AAC(3)-II compromises kanamycin bactericidal activity.
| No | Resistotype | Kanamycin |
Log kill at 24 h (CFU/ml) |
|||
|---|---|---|---|---|---|---|
| MIC | 0.5× MIC | 1× MIC | 2× MIC | 4× MIC | ||
| 8 | Wild-type | 16 | 2.4 | −6.3 | −6.3 | −6.4 |
| 9 | Wild-type | 16 | 2.0 | −6.2 | −6.2 | −6.2 |
| 10 | AAC(3)-IIa | 16 | 2.5 | 2.5 | 1.8 | −1.3 |
| 11 | AAC(3)-IId | 16 | 2.5 | 2.5 | −0.4 | −2.2 |
Boldface red numbers indicate bactericidal activity (≥ 3 log decrease of the bacterial load from the initial inocolum after 24 h of incubation).
WGS indicated a wide diversity in the E. coli strains studied with >70 sequence types (ST) (Table S1), the most prevalent being ST131 (114/488), ST69 (20/488) and ST10 (18/488). Ninety-five isolates displayed either a ST not yet described or no ST could be assigned due to incomplete sequence information. Among the resistance mechanisms identified, five resistance profiles consisted in the production of a single AME [APH(3′), AAC(6′), AAC(3), ANT(2″)] or a single 16S rRNA methyltransferase (RmtB) and eight resistance profiles consisted of various combinations of AMEs and 16S rRNA methyltransferases [AAC(3)/APH(3′); AAC(3)/AAC(6′); ANT(2″)/AAC(6′); ANT(2″)/AAC(3); RmtB/AAC(3); RmtB/AAC(3)/AAC(6′); RmtB/AAC(3)/APH(3′); RmtC/AAC(6′)]. The most prevalent single AMEs were APH(3′) (141/456, 31%), AAC(3) (150/456, 32·9%) and AAC(6′) (53/456, 11·6%).
4. Discussion
The aminoglycoside resistance patterns of 458 E. coli clinical isolates were determined by disc diffusion which, by virtue of its convenience, flexibility, efficiency and low-cost, is probably the most widely used AST method worldwide. The prevalence of AMEs was consistent with data from recent reports from Spain, Poland and Norway, where the most common resistance genes (alone or in combination) are aac(3)-II, aph(3′)-Ia and aac(6′)-Ib-cr [15,31,32]. We established an algorithm (Fig. 1) based on gentamicin, tobramycin and kanamycin ECOFFs, and on an amikacin working separator cut-off, which reliably identified aminoglycoside resistance mechanisms in E. coli (sensitivity of 99.6% and specificity of 97.1%). ARMIA failed to predict resistance only in two cases (0·4%). WGS-based analysis failed to predict resistance in eight isolates (1·6%). Discrepancy analysis (Fig. S2) suggested that most of these vME were likely due to genomic mutations causing broad, low-level aminoglycoside-resistance, such as those altering the respiratory chain [29] or activating efflux pumps [33]. These observations support the notion that WGS cannot replace but rather complement phenotypic testing [34]. In fact, even if WGS becomes as fast and inexpensive as current phenotypic tests, analysis will always be hampered by incomplete data linking genotype and phenotype resulting from complex or yet undescribed mechanisms. Compared to ARMIA and WGS data, EUCAST-based interpretation of in vitro AST results performed poorly with 63 vME. For a summary of the overall performance of the different algorithms see Table 4.
Table 4.
Overall performance of the various algorithms.
| Algorithm | Number of isolates analysed | Wild-type | APH(3′) | AAC(6′)a | AAC(3)b, c | AMEs or AME combinations not affecting AMK | AME combinations affecting AMK and/or RMT | Total | vME |
|---|---|---|---|---|---|---|---|---|---|
| ARMIAd | 488 | Agreement inferred mechanism/genotype | |||||||
| 30/30 (100%) | 139/139 (100%) | 49/51 (96.1%) | 139/143 (97.2%) | 44/44 (100%) | 70/78 (89.7%) | 471/488 (96.5%) | 2 (0.4%) | ||
| WGS-based algorithme | Agreement genotype/ phenotype | 8 (1.6%) | |||||||
| 29/32 (90.6%) | 139/141 (98.6%) | 49/53 (92.5%) | 150/150 (100%) | 37/40 (92.5%) | 72/72 (100%) | 476/488 (97.5%) | |||
| EUCAST-based algorithmf | Agreement inferred mechanism/genotype | 63 (12.9%) | |||||||
| 167/167 (100%) | 50/54 (92.6%) | 184/246 (74.8%) | 17/21 (81.8%) | 418/488 (85.7%) | |||||
AME, aminoglycoside-modifying enzyme; AMK, amikacin; GEN, gentamicin; KAN, kanamycin; RMT: 16S rRNA methyltransferase; TOB, tobramycin; vME, very major error.
In the WGS-based algorithm amikacin test results were disregarded for AAC(6′)-producing isolates.
In the WGS-based algorithm tobramycin and amikacin test results were disregarded for AAC(3)-carrying isolates.
In the WGS-based algorithm the AAC(3)-carrying isolates were included in this category.
For the ARMIA-based analysis, the EUCAST ECOFF for gentamicin (16 mm) and tobramycin (16 mm), the IMM ECOFF for kanamycin (15 mm) and the working separator cut-off for amikacin (22 mm) were used.
For the WGS-based algorithm, the EUCAST ECOFF for gentamicin (16 mm), tobramycin (16 mm) and amikacin (18 mm), and the IMM ECOFF for kanamycin (15 mm) were used.
For the WGS-based algorithm, the EUCAST CBP for gentamicin (17 mm), tobramycin (17 mm) and amikacin (18 mm) were used.
EUCAST guidelines include a number of interpretation rules to infer resistance mechanisms and to review AST results [2,35]. As shown in Fig. S3, the main limitation of EUCAST interpretation of AST results for aminoglycosides and E. coli is the failure to infer the presence of AAC(6′) enzymes when they co-occur with other AMEs [e.g. AAC(3)]. This is due to the fact that amikacin is a ‘poor’ substrate for AAC(6′) due to the presence at position N1 of the 2-deoxystreptamine ring of an L-hydroxyaminobuteroyl amide (L-haba) group that impairs access of this enzyme to the substrate [36]. As a consequence, AAC(6′) enzymes decrease susceptibility to amikacin, but most of the time this does not translate into a change in clinical category, as shown by population analysis of the inhibition zone diameters of strains producing AAC(6′) (Fig. 2). While EUCAST expert rule 12.7 allows to infer the presence an AAC(6′) enzyme from tobramycin resistance, no rules enable the inference of AAC(6′) when combined with other AMEs.
In a proof-of-concept study with a few representative E. coli, we have shown that AAC(6′)-Ib-cr can significantly compromise the in vitro bactericidal activity of amikacin. While at 2× MIC amikacin exerted bactericidal activity against strains devoid of resistance mechanism or harbouring resistance mechanisms not affecting amikacin [APH(3′)-Ia and AAC(3)-IId], amikacin even at 4× MIC was not bactericidal against E. coli producing AAC(6′)-Ib-cr (Fig. S4). These results pair with a study of Klebsiella pneumoniae and together indicate that AAC(6′)-Ib largely abrogates the bactericidal activity of amikacin [37]. Treatment by amikacin in the presence of an AAC(6′)-Ib may then have severe consequences and might result in failure to eradicate the infection. Consistent with this notion, in a rabbit model of experimental endocarditis using a K. pneumoniae producing an AAC(6′)-Ib and categorized as amikacin susceptible, treatment with that drug failed to eradicate the infection [38]. In addition, ‘susceptible’ isolates harbouring a resistance gene may quickly become resistant under therapy due to gene amplification [39]. To solve these issues, we used a working separator cut-off of 22 mm for amikacin that enabled the distinction of AME combinations containing AAC(6′) and affecting amikacin from those not containing AAC(6′) and not affecting amikacin. This allowed to infer the presence of AAC(6′) enzymes in 55/57 strains classified as amikacin susceptible by EUCAST interpretation (Table S7).
Another limitation of EUCAST interpretation was the failure to infer the presence of tobramycin-modifying AAC(3)-II, which led to misclassification of 5/149 AAC(3)-II isolates (3·3%) as tobramycin susceptible. The tobramycin mean inhibition diameters of AAC(3)-II-producing and of wild-type populations are very close, implying that tobramycin is a ‘poor’ substrate for this enzyme (Fig. 2). In ARMIA this issue was addressed by disregarding tobramycin results against gentamicin-resistant isolates [mostly AAC(3)-producers] and by considering these isolates as tobramycin resistant (Fig. 1). This approach allowed to infer the presence of AAC(3)-II enzymes in the five isolates erroneously classified as tobramycin susceptible by EUCAST.
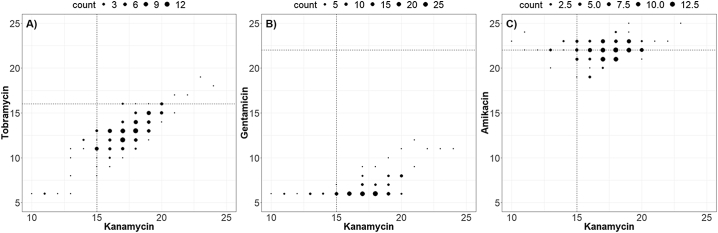
The impact of the AAC(3) enzymes on kanamycin activity has thus far remained unresolved. While Vakulenko and Mobashery reported that only the AAC(3)-III subtype confers resistance to kanamycin [40], Courvalin et al. postulated that all the AAC(3) enzymes confer a kanamycin resistance phenotype [41]. Our population analysis resolved this controversial issue. Analysis of the median kanamycin inhibition zone diameter showed a significant shift for the AAC(3) population as compared to that of the wild-type, indicating a substantial effect of AAC(3) enzymes (mostly of the AAC(3)-II subtype) on kanamycin (Fig. 2). This observation was corroborated by the clear correlation between the inhibition diameters of kanamycin, tobramycin and, in part, gentamicin against the AAC(3)-encoding isolates (Fig. 4A,B). In contrast, no correlation was found between kanamycin and amikacin, which is not a target for AAC(3) (Fig. 4C). In addition, using a few representative isolates, we have shown that AAC(3)-II enzymes affect the bactericidal activity of kanamycin. While at 1× MIC kanamycin was bactericidal against wild-type strains, even at 4× the MIC kanamycin failed to be bactericidal against AAC(3)-II producing E. coli (Fig. S5). Taken together, these data indicate that AAC(3)-harbouring isolates should be considered resistant to kanamycin.
Fig. 4.
AAC(3)-resistotype diameter distributions. Dots correspond to growth inhibition diameters of AAC(3)-harbouring clinical isolates for a) kanamycin and tobramycin, b) kanamycin and gentamicin and c) kanamycin and amikacin. Dot lines indicate EUCAST ECOFFs for gentamicin, tobramycin and amikacin and the IMM ECOFF for kanamycin.
Although the analysis based on ARMIA allows reliable inference of the most clinically-relevant aminoglycoside resistance mechanisms by using only four antibiotics, this study presents some limitations. First, in most routine diagnostic laboratories AST of pathogens is usually performed with gentamicin and amikacin, two of the four aminoglycosides required for ARMIA. Implementation of two additional antibiotic discs will thus be required to ensure reliable identification of all aminoglycoside resistance mechanisms. Second, some AMEs [i.e. ANT(2″), ANT(4′), AAC(3)-I] were either not included in the study or were poorly represented. Third, this was a single-centre study and all the E. coli clinical isolates were collected in Zurich or its surroundings. Although single-centre studies are better suited for method validation, the use of regionally restricted clinical isolates might have limited the genetic diversity of the resistotypes. Fourth, the recently FDA-approved plazomicin was not included in the algorithm, since it was not available at the time of the study.
Enterobacterales have nearly identical ECOFFs and very similar susceptibility profiles towards aminoglycosides [3]. As a result, EUCAST recommends the same CBPs for the Enterobacterales species, which implies that ARMIA may be used for interpretation of aminoglycoside AST for all the species of the Enterobacterales order.
In conclusion, we have demonstrated that interpretative reading of phenotypic aminoglycoside susceptibility data can generate reliable information on the underlying resistance mechanisms, with efficiency comparable to that of WGS. The algorithm developed provides a coherent phenotype-genotype interpretation system and may be integrated in the existing EUCAST expert rule set. The study illustrates the impact of WGS and population analysis on designing interpretative AST rules for the diagnostic laboratory and highlights the importance of accurate and reliable inference of resistance mechanisms for selection of the most appropriate antibiotic regimen.
Contributors
The study was designed by E.C. Böttger and P. Courvalin. Data were produced by M. Marchesi, S. Mancini, C. Quiblier, F. Imkamp, K. Wagner and E. Bodendörfer. Data were analysed by S. Mancini, M. Marchesi, E.C. Böttger, P. Courvalin and F. Imkamp. In the initial phase of the study P. Keller was involved in the analysis of the data. Data were interpreted by E.C. Böttger, P. Courvalin and S. Mancini. The manuscript was written by S. Mancini, E.C. Böttger and P. Courvalin.
Funding
This work was supported by the University of Zurich. The funder did not have any role in the study design, data collection, data analysis, interpretation and writing of the report.
Acknowledgments
Acknowledgments
We thank the technicians of the IMM and the University of Zurich for continuous support. We are grateful to N Blöchliger for help in various phases of the study, J Allgäuer for performing AST and V Pereira Pires for technical assistance. We thank J Kalinowski (CeBiTec, Bielefeld University, Germany) for WGS data and S Elmer for technical support.
Declaration of Competing Interest
All authors: No conflicts of interest to declare.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.ebiom.2019.07.020.
Appendix A. Supplementary data
Supplementary material 1
Supplementary material 2
References
- 1.Courvalin P. Interpretive reading of antimicrobial susceptibility tests. ASM News. 1992;58:368–375. [Google Scholar]
- 2.European Committee on Antimicrobial Susceptibility Testing EUCAST guidelines for the detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance. 2018. http://www.eucast.org/resistance_mechanisms/
- 3.European Committee on Antimicrobial Susceptibility Testing . EUCAST; Växjö, Sweden: 2018. MIC and zone distributions and ECOFFs.http://mic.eucast.org/Eucast2/ [Google Scholar]
- 4.Balloux F., Bronstad Brynildsrud O., van Dorp L. From theory to practice: translating whole-genome sequencing (WGS) into the clinic. Trends Microbiol. 2018;26(12):1035–1048. doi: 10.1016/j.tim.2018.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ellington M.J., Ekelund O., Aarestrup F.M. The role of whole genome sequencing in antimicrobial susceptibility testing of bacteria: report from the EUCAST subcommittee. Clin Microbiol Infect. 2017;23(1):2–22. doi: 10.1016/j.cmi.2016.11.012. [DOI] [PubMed] [Google Scholar]
- 6.CRyPTIC C, The GP, Allix-Beguec C. Prediction of susceptibility to first-line tuberculosis drugs by DNA sequencing. N Engl J Med. 2018;379(15):1403–1415. doi: 10.1056/NEJMoa1800474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Deggim-Messmer V., Bloemberg G.V., Ritter C. Diagnostic molecular mycobacteriology in regions with low tuberculosis endemicity: combining real-time PCR assays for detection of multiple mycobacterial pathogens with line probe assays for identification of resistance mutations. EBioMedicine. 2016;9:228–237. doi: 10.1016/j.ebiom.2016.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gygli S.M., Keller P.M., Ballif M. Whole-genome sequencing for drug resistance profile prediction in Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2019;63(4) doi: 10.1128/AAC.02175-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tangden T. Combination antibiotic therapy for multidrug-resistant Gram-negative bacteria. Ups J Med Sci. 2014;119(2):149–153. doi: 10.3109/03009734.2014.899279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ramirez M.S., Tolmasky M.E. Aminoglycoside modifying enzymes. Drug Resist Updat. 2010;13(6):151–171. doi: 10.1016/j.drup.2010.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Davis M.A., Besser T.E., Orfe L.H. Genotypic-phenotypic discrepancies between antibiotic resistance characteristics of Escherichia coli isolates from calves in management settings with high and low antibiotic use. Appl Environ Microbiol. 2011;77(10):3293–3299. doi: 10.1128/AEM.02588-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fernandez-Martinez M., Miro E., Ortega A. Molecular identification of aminoglycoside-modifying enzymes in clinical isolates of Escherichia coli resistant to amoxicillin/clavulanic acid isolated in Spain. Int J Antimicrob Agents. 2015;46(2):157–163. doi: 10.1016/j.ijantimicag.2015.03.008. [DOI] [PubMed] [Google Scholar]
- 13.Miro E., Grunbaum F., Gomez L. Characterization of aminoglycoside-modifying enzymes in Enterobacteriaceae clinical strains and characterization of the plasmids implicated in their diffusion. Microb Drug Resist. 2013;19(2):94–99. doi: 10.1089/mdr.2012.0125. [DOI] [PubMed] [Google Scholar]
- 14.Davis B.D. Mechanism of bactericidal action of aminoglycosides. Microbiol Rev. 1987;51(3):341–350. doi: 10.1128/mr.51.3.341-350.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ojdana D., Sienko A., Sacha P. Genetic basis of enzymatic resistance of E. coli to aminoglycosides. Adv Med Sci. 2018;63(1):9–13. doi: 10.1016/j.advms.2017.05.004. [DOI] [PubMed] [Google Scholar]
- 16.van de Klundert J.A., Vliegenthart J.S., van Doorn E., Bongaerts G.P., Molendijk L., Mouton R.P. A simple method for the identification of aminoglycoside-modifying enzymes. J Antimicrob Chemother. 1984;14(4):339–348. doi: 10.1093/jac/14.4.339. [DOI] [PubMed] [Google Scholar]
- 17.Bossuyt P.M., Reitsma J.B., Bruns D.E. STARD 2015: an updated list of essential items for reporting diagnostic accuracy studies. BMJ. 2015;351 doi: 10.1136/bmj.h5527. h5527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.European Committee on Antimicrobial Susceptibility Testing . EUCAST; Växjö, Sweden: 2010. Setting breakpoints for new agents standard operating procedure 11.http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/EUCAST_SOPs/EUCAST_SOP_1._1_Setting_breakpoints_new_agents_1_June_3.pdf [Google Scholar]
- 19.Galimand M., Sabtcheva S., Courvalin P., Lambert T. Worldwide disseminated armA aminoglycoside resistance methylase gene is borne by composite transposon Tn1548. Antimicrob Agents Chemother. 2005;49(7):2949–2953. doi: 10.1128/AAC.49.7.2949-2953.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.European Committee on Antimicrobial Susceptibility Testing . EUCAST; Växjö, Sweden: 2018. EUCAST disk diffusion method for antimicrobial susceptibility testing, version 60; p. 2017.http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Disk_test_documents/Version_5/Manual_v_6.0_EUCAST_Disk_Test_final.pdf [Google Scholar]
- 21.Hombach M., Zbinden R., Bottger E.C. Standardisation of disk diffusion results for antibiotic susceptibility testing using the sirscan automated zone reader. BMC Microbiol. 2013;13:225. doi: 10.1186/1471-2180-13-225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.European Committee on Antimicrobial Susceptibility Testing EUCAST reading guide for broth microdilution. 2019. http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Disk_test_documents/2019_manuals/Reading_guide_BMD_v_1.0_.pdf
- 23.Hunt M., Mather A.E., Sanchez-Buso L. ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Microb Genom. 2017;3(10) doi: 10.1099/mgen.0.000131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Gupta S.K., Padmanabhan B.R., Diene S.M. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob Agents Chemother. 2014;58(1):212–220. doi: 10.1128/AAC.01310-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.McArthur A.G., Waglechner N., Nizam F. The comprehensive antibiotic resistance database. Antimicrob Agents Chemother. 2013;57(7):3348–3357. doi: 10.1128/AAC.00419-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wirth T., Falush D., Lan R. Sex and virulence in Escherichia coli: an evolutionary perspective. Mol Microbiol. 2006;60(5):1136–1151. doi: 10.1111/j.1365-2958.2006.05172.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.R Core Team R: A language and environment for statistical computing. R foundation for statistical computing. 2015. https://www.R-project.org/ Vienna, Austria.
- 28.Witchitz J.L. Plasmid-mediated gentamicin resistance not associated with kanamycin resistance in Enterobacteriaceae. J Antibiot (Tokyo) 1972;25(10):622–624. doi: 10.7164/antibiotics.25.622. [DOI] [PubMed] [Google Scholar]
- 29.Hancock R.E., Farmer S.W., Li Z.S., Poole K. Interaction of aminoglycosides with the outer membranes and purified lipopolysaccharide and OmpF porin of Escherichia coli. Antimicrob Agents Chemother. 1991;35(7):1309–1314. doi: 10.1128/aac.35.7.1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Robicsek A., Strahilevitz J., Jacoby G.A. Fluoroquinolone-modifying enzyme: a new adaptation of a common aminoglycoside acetyltransferase. Nat Med. 2006;12(1):83–88. doi: 10.1038/nm1347. [DOI] [PubMed] [Google Scholar]
- 31.Fernandez-Martinez M., Ruiz Del Castillo B., Lecea-Cuello M.J. Prevalence of aminoglycoside-modifying enzymes in Escherichia coli and Klebsiella pneumoniae producing extended spectrum beta-lactamases collected in two multicenter studies in Spain. Microb Drug Resist. 2018;24(4):367–376. doi: 10.1089/mdr.2017.0102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Haldorsen B.C., Simonsen G.S., Sundsfjord A., Samuelsen O., Norwegian Study Group on Aminoglycoside R Increased prevalence of aminoglycoside resistance in clinical isolates of Escherichia coli and Klebsiella spp. in Norway is associated with the acquisition of AAC(3)-II and AAC(6′)-Ib. Diagn Microbiol Infect Dis. 2014;78(1):66–69. doi: 10.1016/j.diagmicrobio.2013.10.001. [DOI] [PubMed] [Google Scholar]
- 33.Rosenberg E.Y., Ma D., Nikaido H. AcrD of Escherichia coli is an aminoglycoside efflux pump. J Bacteriol. 2000;182(1756):1754–1756. doi: 10.1128/jb.182.6.1754-1756.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Koser C.U., Ellington M.J., Cartwright E.J. Routine use of microbial whole genome sequencing in diagnostic and public health microbiology. PLoS Pathog. 2012;8(8) doi: 10.1371/journal.ppat.1002824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.European Committee on Antimicrobial Susceptibility Testing EUCAST expert rules in antimicrobial susceptibility testing 2018. 2019. http://www.eucast.org/expert_rules_and_intrinsic_resistance/
- 36.Ramirez M.S., Tolmasky M.E. Amikacin: uses, resistance, and prospects for inhibition. Molecules. 2017;22(12) doi: 10.3390/molecules22122267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Bremmer D.N., Clancy C.J., Press E.G. KPC-producing Klebsiella pneumoniae strains that harbor AAC(6′)-Ib exhibit intermediate resistance to amikacin. Antimicrob Agents Chemother. 2014;58(12):7597–7600. doi: 10.1128/AAC.03831-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Caulin E., Coutrot A., Carbon C., Collatz E. Resistance to amikacin and isepamicin in rabbits with experimental endocarditis of an aac(6′)-Ib-bearing strain of Klebsiella pneumoniae susceptible in vitro. Antimicrob Agents Chemother. 1996;40(12):2848–2853. doi: 10.1128/aac.40.12.2848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.McGann P., Courvalin P., Snesrud E. Amplification of aminoglycoside resistance gene aphA1 in Acinetobacter baumannii results in tobramycin therapy failure. MBio. 2014;5(2) doi: 10.1128/mBio.00915-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Vakulenko S.B., Mobashery S. Versatility of aminoglycosides and prospects for their future. Clin Microbiol Rev. 2003;16(3):430–450. doi: 10.1128/CMR.16.3.430-450.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Courvalin P., Leclercq L., Rice L.B. ASM Press; Washington, D.C.: 2010. Antibiogram. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary material 1
Supplementary material 2