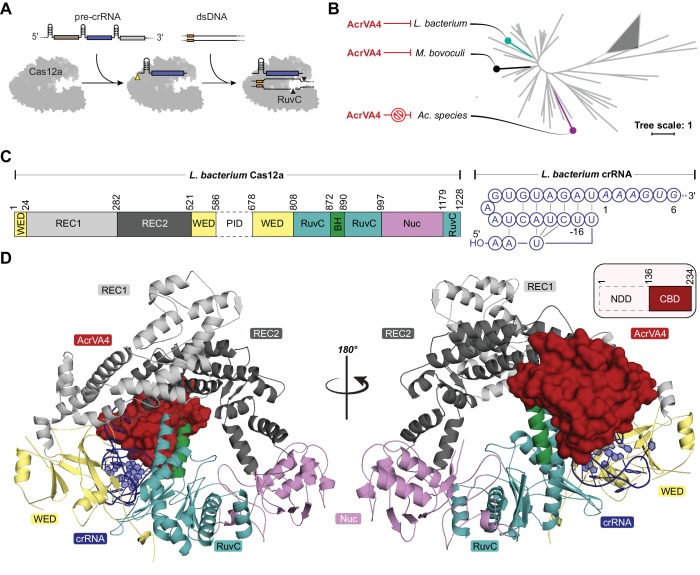
Figure 1. Overall structure of the LbCas12a-crRNA-AcrVA4 complex.
(A) Schematic representation of Cas12a activity. (B) Unrooted maximum likelihood phylogenetic tree of Type V-A CRISPR-Cas12a. Species known to be susceptible or unsusceptible to phage-derived AcrVA4 are highlighted. The triangle denotes collapsed branches of Cas12b-e. (C) Schematic representation of LbCas12a and the mature crRNA modeled within the cryo-EM structures. (D) Two views of the LbCas12a-crRNA complex (cartoon) bound to AcrVA4 (surface) shown related by a 180° rotation. The color scheme for the crRNA, LbCas12a, and AcrVA4 in panels B, C, and D are used throughout the manuscript.