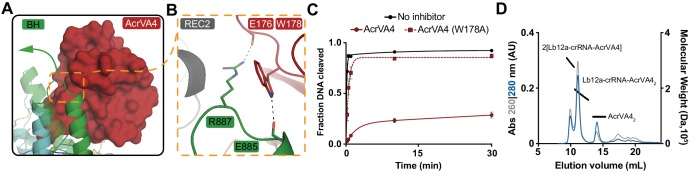
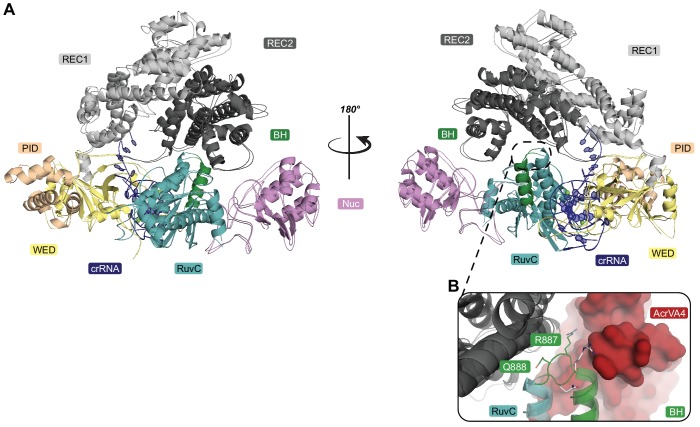
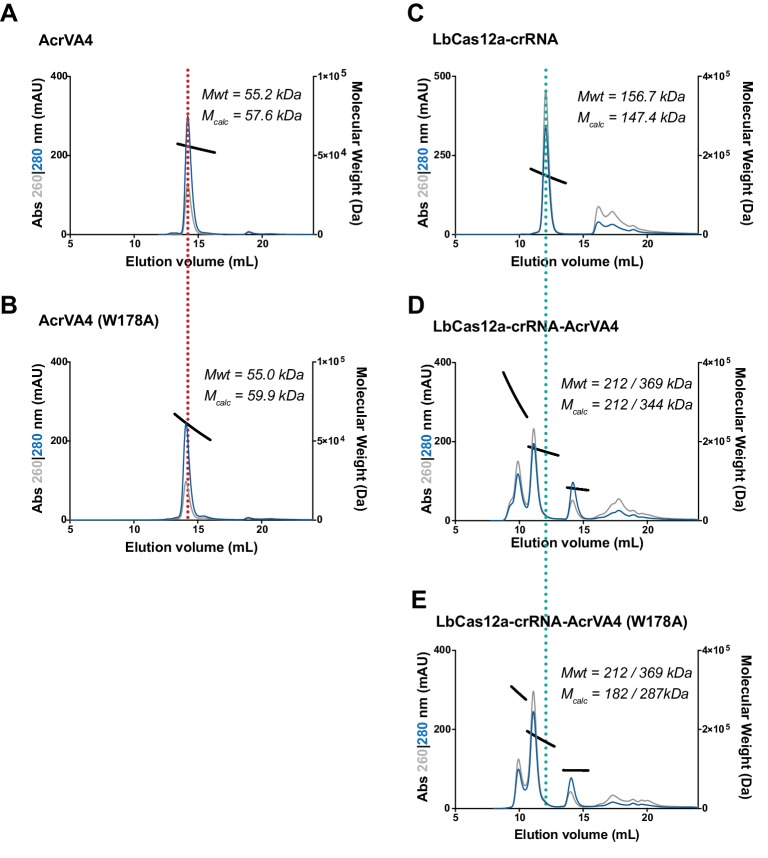
Figure 4. AcrVA4 locks the Cas12a bridge-helix to prevent DNA binding.
(A) AcrVA4 (surface) is shown braced against the bridge-helix (BH, green, cartoon) near the RuvC (teal, cartoon). The conformation of the LbCas12a bridge-helix when bound to DNA is shown semi-transparent with a green arrow denoting the direction of helix motion upon DNA binding. (B) Detailed atomistic view of AcrVA4 (red) recognition of the Cas12a bridge-helix (green). (C) LbCas12a dsDNA cis-cleavage over time measured under single-turnover conditions in the presence or absence of AcrVA4 containing alanine substitutions (mean ∓ s.d., n = 3 independent measurements). Two-phase exponential decay experimental fits are shown as solid or dashed lines. (D) Size-exclusion chromatography coupled di-angle light scattering (SEC-DALS) trace for AcrVA4 W178A in the presence of LbCas12a-crRNA. The absorbance at 280 nm (blue) and 260 nm (gray) are shown (left axis) with the linear region for the mass estimate corresponding to the relevant peaks (black lines, right axis).