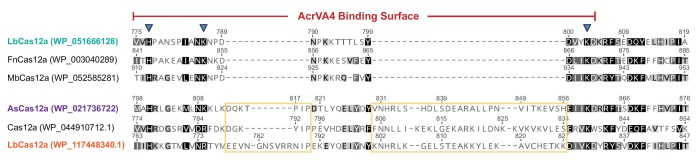
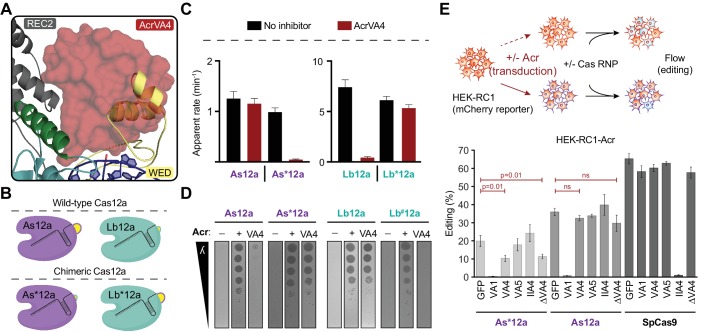
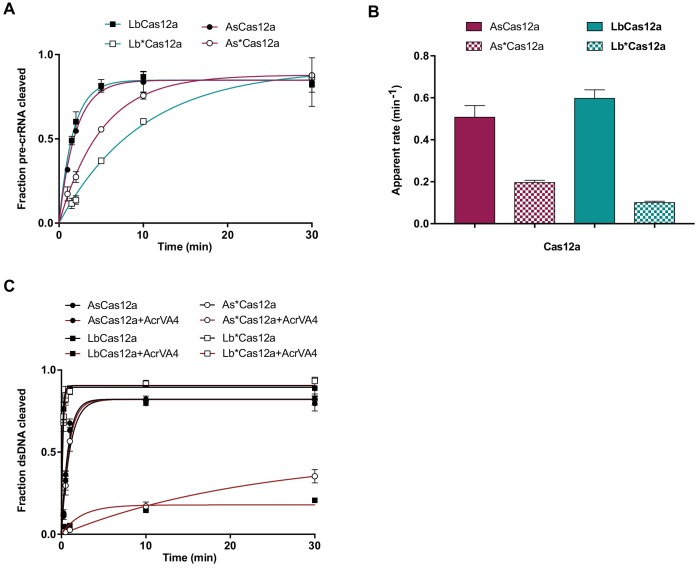
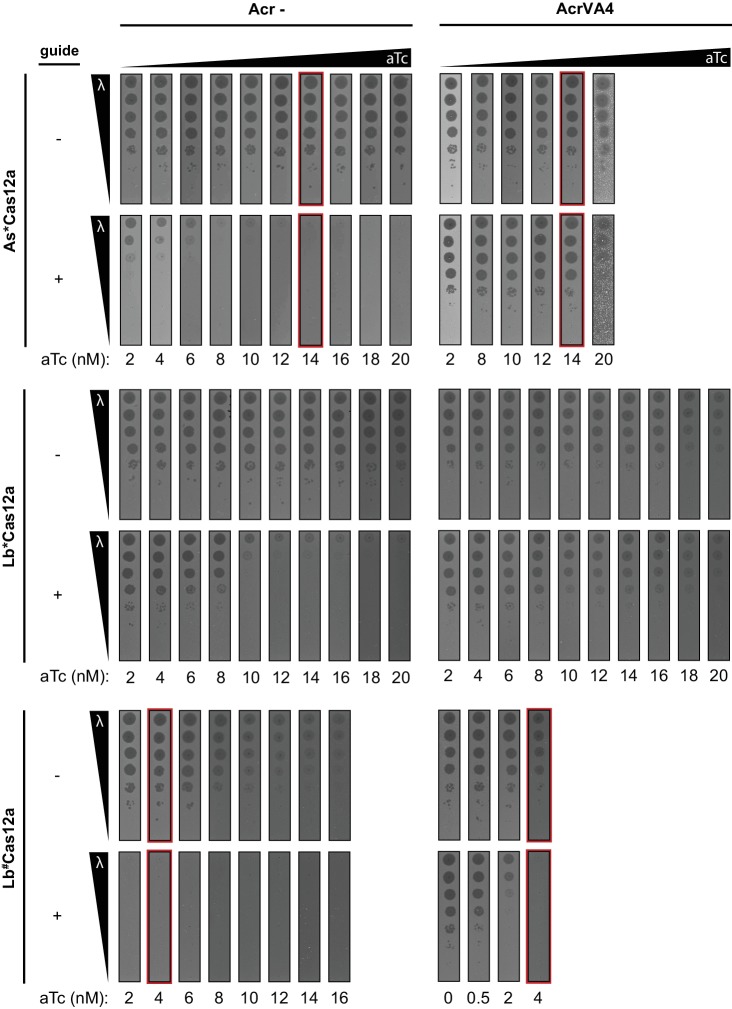
Figure 6. AsCas12a evades AcrVA4 by concealing its pre-crRNA processing nuclease.
(A) The C-terminal binding domain of AcrVA4 (surface) is shown superposed on the AsCas12a structure where it clashes with the WED domain (yellow, cartoon). (B) Schematic representation of the wild-type (Lb and As) and engineered chimeric (Lb* and As*) Cas12a constructs. (C) Bar-graph illustrating the apparent rate of wild-type or chimeric Cas12a-mediated dsDNA cis-cleavage under single-turnover conditions in the presence or absence of AcrVA4 (mean ∓ s.d., n = 3 independent measurements). Single-phase exponential fits to the cleavage kinetics from which the rate was derived can be found in Figure 6—figure supplement 2C. (D) Phage plaque assays to determine susceptibility of chimeric Cas12a to AcrVA4 in E. coli. Ten-fold serial dilutions of heat-inducible phage lambda spotted on lawns of E. coli strains expressing lambda-targeting guide, wild-type or chimeric (*/#) Cas12a, and AcrVA4 or the indicated control anti-CRISPR protein. Images shown are representative of the effect seen in replicates (n = 3 independent measurements). (E) CRISPR-Cas12a inhibition specificity in human cells. Schematic (top panel) showing human cells stably expressing a fluorescence reporter and doxycycline-inducible anti-CRISPR (Acr) constructs. Acr expression blocks genome editing upon transfection of susceptible Cas ribonucleoprotein (RNP) complexes, quantifiable by flow cytometry of mCherry fluorescence. Assessment of editing efficiency in HEK-RC1 reporter cells (bottom panel) expressing GFP or GFP-Acr polycistronic constructs (AcrVA1, AcrVA4, AcrVA5, AcrIIA4, AcrVA4 Δ1–134) and transfected with various Cas9 and Cas12a RNPs targeting the reporter. Note, in contrast to wild-type AsCas12a (As12a), editing by the AsCas12a-chimera (As*12a) was moderately susceptible to AcrVA4 and AcrVA4 Δ1–134 inhibition.