Figure 3.
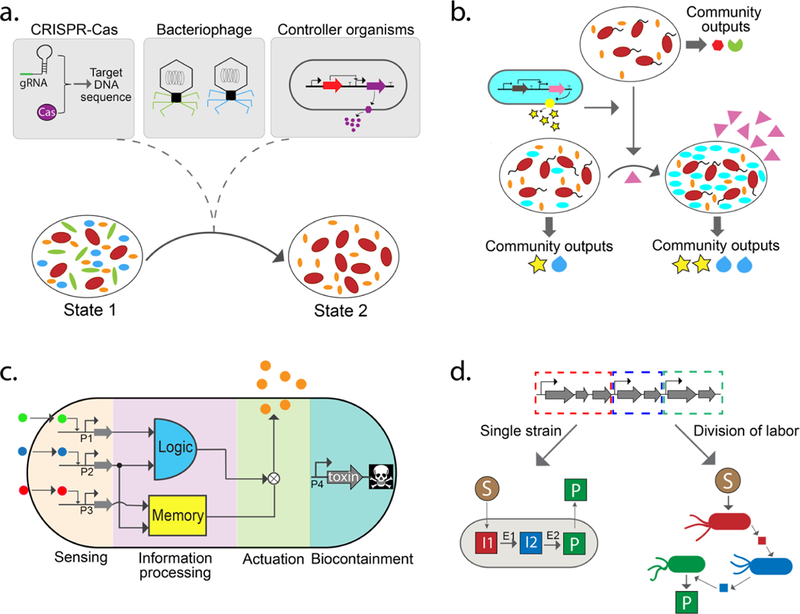
Strategies for engineering natural and synthetic communities. (a) Precise inhibition of specific organisms in natural communities via (i) CRISPR-guided nucleases that target specific DNA sequences, (ii) engineered bacteriophages, and (iii) microbes designed for production of antimicrobial compounds. (b) Introduction of an engineered organism (cyan) into a microbial community. The engineered organism can stably integrate into the community and alters community-level outputs (represented by yellow stars and blue teardrops). The abundance of the new species could be controlled through niche engineering, wherein a non-native organism is engineered to utilize a unique substrate (pink triangles) that cannot be utilized by other members of the community. (c) Design of cellular control organisms using engineered biomolecular networks to perform novel functions in complex environments. Functional modules include environmental sensors, information processing (e.g., memory and logic circuits), actuators (e.g., antimicrobial compounds), and growth control circuits for biocontainment. (d) Construction of synthetic communities from the bottom up to perform desired functions through division of labor (DOL). Top: A single biochemical pathway can be conceptually decomposed into three subpathways (red, blue, and green). Left arrow: The biochemical pathway for synthesizing a target molecule can be introduced into a single cell population. Right arrow: The biochemical pathway is partitioned among different organisms, reducing the metabolic burden on individual populations but requiring stable coexistence as a function of time. S, substrate; P, product; I, intermediate; E, enzyme.
