Abstract
Although DNA transposons often generated internal deleted derivatives such as miniature inverted-repeat transposable elements, short internally deleted elements (SIDEs) derived from nonlong terminal-repeat retrotransposons are rare. Here, we found a novel SIDE, named Persaeus, that originated from the chicken repeat 1 (CR1) retrotransposon Zenon and it has been found widespread in Lepidoptera insects. Our findings suggested that Persaeus and the partner Zenon have experienced a transposition burst in their host genomes and the copy number of Persaeus and Zenon in assayed genomes are significantly correlated. Accordingly, the activity though age analysis indicated that the replication wave of Persaeus coincided with that of Zenon. Phylogenetic analyses suggested that Persaeus may have evolved at least four times independently, and that it has been vertically transferred into its host genomes. Together, our results provide new insights into the evolution dynamics of SIDEs and its partner non-LTRs.
Keywords: chicken repeat 1 (CR1), transposable elements evolutionary dynamics, long interspersed element (LINE), Lepidoptera, short internally deleted element (SIDE), vertical inheritance
Introduction
The eukaryotic genome is composed by a wide diversity of transposable elements (TE), some autonomous (i.e., coding for the enzymatic machinery necessary for replication and reintegration) and some others nonautonomous (i.e., dependent on autonomous-encoded enzymes for replication and reintegration) (Chénais et al. 2012). Among nonautonomous elements, there are the short interspersed elements (SINEs) that are nucleotide (nt) sequence made by different modules (head, body, and tail) with different origins (Luchetti and Mantovani 2013). Other kind nonautonomous elements are internally deleted copies of autonomous elements. Miniature inverted-repeat transposable elements (Feschotte and Pritham 2007) and terminal-repeat retrotransposons in miniature (Gao et al. 2016) are widespread elements, derived from internal deletions of autonomous DNA transposons and long-terminal-repeat retrotransposons (LTR), respectively. On the contrary, short internally deleted elements (SIDEs) originated from non-LTR elements seems to be rare, being only found in fruit flies, in the mosquito Anopheles gambiae and in the protozoan Trypanosoma brucei (Kimmel et al. 1987; Biedler and Tu 2003; Eickbush and Eickbush 2012).
Chicken repeat 1 (CR1) elements are non-LTRs, long interspersed elements (LINEs) and were the first TE found in the chicken genome about three decades ago (Stumph et al. 1981, 1984). CR1 replicates through a “copy-and-paste” mechanism and, usually, shows two open reading frames (ORFs) coding for a Gag-like protein, which has a zinc finger motif, and a Pol-like protein, which has endonuclease and reverse transcriptase (RT) domains (Burch et al. 1993; Haas et al. 1997; Kajikawa et al. 1997). Compared with L1 LINE, 5′-UTR of CR1 elements are more frequently truncated, which imply a lower processivity of its transcription (Hillier et al. 2004).
CR1 elements are the most abundant TE families in the genomes of birds (Hillier et al. 2004; Warren et al. 2010), crocodilians (Green et al. 2014), snakes (Castoe 2013), and turtles (Shaffer et al. 2013) and are composed by a great diversity that existed from the era of the common ancestor of amniotes (Suh et al. 2014). CR1 elements are also the only active TEs throughout the evolution of birds and, thus, have been widely served as genetic markers (Kaiser et al. 2006; Haddrath and Baker 2012; Liu et al. 2012; Baker et al. 2014). However, the evolutionary history and dynamics of CR1 elements in insects remain largely unknown. So far, CR1 have been found in a few insects, namely some flies (Kapitonov and Jurka 2003; Thompson et al. 2009), the mosquito A. gambiae (Biedler and Tu 2003) and some Lepidoptera (butterflies and moths) species (Novikova et al. 2007), where it may show even only a single ORF encoding endonuclease and RT domains. To our best knowledge, there was only one documented example of SIDEs originated from CR1 elements (Biedler and Tu 2003).
In this study, we report on the finding of a novel SIDE, derived from a CR1 element, isolated from the genome of Lepidoptera insects. Obtained results suggested that this SIDE as well as its partner Zenon have been highly active during the evolution of some Lepidoptera superfamilies and that the SIDE may have evolved multiple times, independently. Moreover, although widespread among Lepidoptera, our results suggest a vertical inheritance at least at lower taxonomic level. Overall, we concluded that SIDE and Zenon reported here might provide a good system to study the dynamics of emergence of SIDEs and their interaction with the partner LINE.
Materials and Methods
Animal Materials
Dazao, a strain of the silkworm B. mori, was obtained from the State Key Laboratory of Silkworm Genome Biology (China). Antheraea pernyi and A. yamamai were collected from Heilongjiang province (China) and Changbai Mountain (Jilin province, China), respectively. Rhodnius prolixus was kindly provided by Dr Ricardo Nascimento Araujo (Laboratório de Fisiologia de Insetos Hematófagos, Brazil). Samia insularis, Samia luzonica, Samia cynthia ricini, Amathuxidia amythaon and Caligo eurilochus was purchased from Shanghai Qiuyu Biotechnology Co., Ltd (China). Then, we extracted their total DNAs using TIANamp Genomic DNA Kit (TIANGEN).
PCR, Cloning, and Sequencing
We designed a pair of specific primers (Forward: 5′-GAG CCG ATT GTT GAA GCG GAA AAA G-3′; Reverse: 5′-TGG CCT TGA TAG CGT TGT TCA AAA T-3′) of Garfield_BM (Zhang et al. 2014) using its internal sequence to determine its distribution in some insects. PCR was performed with an initial denaturation step of 4 min at 95 °C followed by 30 cycles of 40 s at 95 °C, 40 s at 58 °C, and 2 m at 72 °C. Then, purified PCR products were cloned into PMD-19 cloning vector (TaKaRa). One or two random clones of each species were selected and sequenced.
Sequence Analyses
Two SIDEs search strategies have been implemented. In the first, SIDEs were found in published Lepidoptera genomes by BLASTing the A. pernyi SIDE sequence with the blastn algorithm and e-value >10−10. In the second, Zenon was first found by means of tblastn algorithm (e-value >10−5) of BLAST search using the RT domain as query sequence; once characterized the Zenon nt sequence, the 5′ and the 3′ end where manually joined and used to BLAST search as described above. When the search gave significant positive hits, the first full-length 50 hits were used to build a majority rule consensus sequence. This consensus sequence was, then, used to perform an exhaustive search on relative genomes using the same BLAST search parameters. All positive hits were used to build a new, final SIDE consensus sequence for each genome. In addition to genomes scan, also the nonredundant nt, ESTs, and TSA NCBI databases (accessed on May 2019) were probed with all consensus sequences in order to find further SIDE copies.
The search for partner LINE Zenon elements was performed following the same procedure. The only exception was that in some instances no full-length copies were retrieved: When possible, the complete Zenon sequence was reconstructed by manually aligning BLAST hit regions and recognizing the element borders. In some instance, we were unable to reconstruct the full-length sequence, so that those elements were no further considered. All obtained consensus sequences were, then, validate by checking the presence of ORF translating in an RT domain. SIDE and partner LINE copy number determination and activity through age analysis have been carried out on genomes using RepeatMasker v. 4.0 (Smit et al. 2013–2015). However, because the homology between the SIDE and the LINE could determine that consensus sequences mask each other copies we decided to exclude fragments long <160 bp (250 bp in the case of Leptidea sinapis): This allowed to recover fragment unambiguously belonging to the SIDE or to the LINE (fig. 1). Moreover, to further refine the copy number estimation, adjacent fragments were merged into single hits using the script Onecodetofindthemall.pl (Bailly-Bechet et al. 2014).
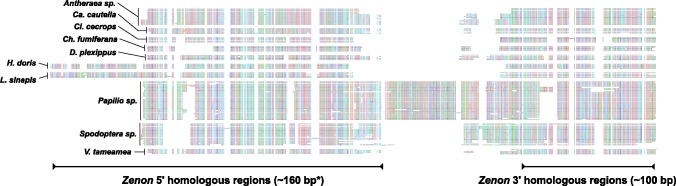
Fig. 1.
—Schematic view of Persaeus sequences (five copies per species) with indication of Zenon homologous regions. Approximate length of homologous regions is also reported (* H. doris and L. sinapis homologous 5′ end is ∼250 bp).
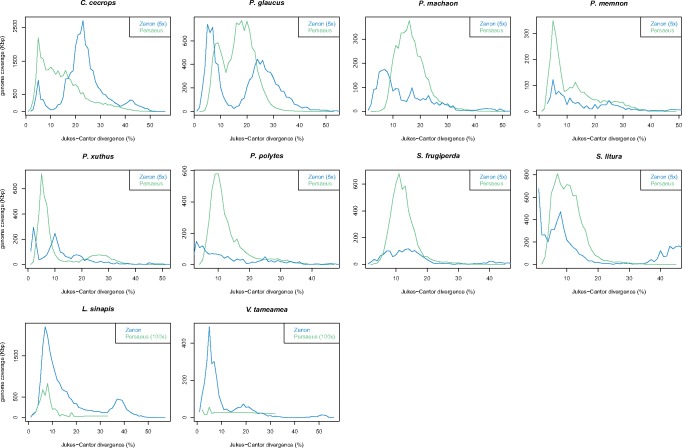
In the activity through age analysis, the relative repeat abundances are plotted against the Jukes–Cantor genetic divergence (which takes into account also multiple substitutions) of each repeat copy versus the consensus sequence of its family. The less divergent copies are the most recently transposed, and the most divergent are those whose replication occurred far in the past.
Phylogenetic analyses were carried out through maximum likelihood and Bayesian inference. Maximum likelihood was performed with RAxML v.8 (Stamatakis 2014) using the GTR+G substitution model (Parsaeus and Zenon nt data sets) or rtREV+G model (Zenon RT amino acid data set) and 100 rapid bootstrap replicates. Bayesian inference was done using MrBayes v. 3.2 (Ronquist et al. 2012), with the same models as above: 2 independent runs searched for 106 generation and trees were sampled every 100. Convergence of the two runs was reached when the average variance of split frequencies <0.01 and Potential Scale Reduction Factor approached ∼1.0. The final Bayesian consensus tree was obtained after a conservative burnin=25%.
Results
Identification of a Novel SIDE and Its Partner LINE
A survey on the distribution of a Chapaev transposon named Garfield identified in our previous study (Zhang et al. 2014) in some insects was performed using polymerase chain reaction (PCR): One PCR amplification band obtained from the Chinese tussar moth Antheraea pernyi was ∼350 bp longer than the expected band size (supplementary fig. S1, Supplementary Material online). After cloning and sequencing, we found that Garfield from the Chinese tussar moth had an additional insertion of 352 bp. This insertion exhibited a poly-(A) 3′ end and seemed to be flanked by a (T)6 target site duplication (supplementary fig. S2, Supplementary Material online). A homology search in the Repbase Update database (Jurka 2000; last accessed October 2018) evidenced that the full length of this insertion shared ∼70% of nt sequence identity with CR1 autonomous elements, named Zenon, from two lepidopteran species: Heliconius melpomene and Papilio xuthus. More in detail, a sequence comparison indicated that homologous regions are at the 5′ end, overlapping the 5′-UTR and the beginning of the Zenon ORF, and at the 3′ end, encompassing the end of the ORF and the whole 3′-UTR of the Zenon elements including the poly-(A) tail (supplementary fig. S3, Supplementary Material online). The insertion does not show any homology with those of tRNA, 5S rRNA, or 7S rRNA genes and lacks an RNA pol III promoter, which are two major characteristics that distinguish SINEs from other nonautonomous transposons (Luchetti and Mantovani 2013). Therefore, this suggested that the insertion found in the Garfield element from the Chinese tussar moth is, actually, an SIDE derived from an internal deletion of the Zenon element. This novel SIDE has been named Persaeus, as he was the favorite disciple of the Greek philosopher Zenon of Citium.
Taxonomic Distribution of Persaeus and Zenon
We investigated the distribution of the SIDE Persaeus and its partner LINE Zenon in other genomes available at the National Center for Biotechnology Information (NCBI; last accessed June 2019), including the nonredundant nt, expressed sequence tags (EST), and transcriptome sequences assembly (TSA). We found that Persaeus was present in the genome of 21 Lepidoptera species belonging to the Bombycoidea, Pyraloidea, Papilionoidea, Tortricoidea, Noctuoidea superfamilies (table 1). The copy number ranged from 12 in Vanessa tameamea (Papilionoidea) to 115,283 in Calycopis cecrops (Papilionoidea), covering up to the 3.68% of the genome (supplementary table S1, Supplementary Material online). We found Zenon in the same also present in additional six species, two of which belonging to further superfamilies: Gelechioidea and Hesperioidea. The copy number varied from 245 in Danaus chrysippus (Papilionoidea) to 40,029 in in Leptidea sinapis (Papilionoidea; table 1); they cover up to the 3.45% of the genome of Leptidea sinapis (supplementary table S1, Supplementary Material online). No positive hits were found outside Lepidoptera in any peered database.
Table 1.
Detailed Information of Persaeus SIDEs and the Associated LINE Zenon in This Study
| Species | Taxonomy (Superfamily) | Persaeus | Persaeus Copy Number | Persaeus Length (bp) | Zenon | Zenon Copy Number | Zenon Length (bp) | Database/Genome Assembly Acc. no. |
|---|---|---|---|---|---|---|---|---|
| Antheraea assama | Bombycoidea | ✓ | 271 | ✓ | 3,315 | TSA | ||
| A. pernyi | Bombycoidea | ✓ | 317 | ✓ | 3,316 | TSA | ||
| A. yamamai | Bombycoidea | ✓ | 302 | TSA | ||||
| Bombyx mandarina | Bombycoidea | ✓ | 5,963 | 3,534 | GCA_003987935.1 | |||
| Cadra cautella | Pyraloidea | ✓ | 316 | TSA | ||||
| Calephelis nemesis | Papilionoidea | ✓ | 8,938 | 3,461 | GCA_002245505.1 | |||
| Calycopis cecrops | Papilionoidea | ✓ | 115,283 | 263 | ✓ | 12,324 | 3,386 | GCA_001625245.1 |
| Choristoneura fumiferana | Tortricoidea | ✓ | 274 | EST | ||||
| Danaus chrysippus | Papilionoidea | ✓ | 245 | 3,352 | GCA_004959915.1 | |||
| Danaus plexippus | Papilionoidea | ✓ | 3,019 | 304 | GCA_000235995.2 | |||
| Heliconius melpomene | Papilionoidea | ✓ | 1,461 | 3,392 | GCA_000313835.2 | |||
| H. numata | Papilionoidea | ✓ | 1,115 | 3,396 | GCA_900068715.1 | |||
| H. doris | Papilionoidea | ✓ | 821 | 341 | GCA_900068325.1 | |||
| Hyposmocoma kahamanoa | Gelechioidea | ✓ | 6,419 | 3,064 | GCA_003589595.1 | |||
| Leptidea sinapis | Papilionoidea | ✓ | 158 | 366 | ✓ | 40,029 | 3,366 | GCA_900199415.1 |
| Megathymus ursus | Hesperioidea | ✓ | 14,562 | 3,741 | GCA_003671415.1 | |||
| Papilio dardanus | Papilionoidea | ✓ | 382 | nt | ||||
| P. glaucus | Papilionoidea | ✓ | 37,447 | 386 | ✓ | 5,516 | 3,340 | GCA_000931545.1 |
| P. machaon | Papilionoidea | ✓ | 15,080 | 386 | ✓ | 1,502 | 3,282 | GCA_001298355.1 |
| P. memnon | Papilionoidea | ✓ | 8,993 | 381 | ✓ | 752 | 3,300 | GCA_003118335.3 |
| P. polytes | Papilionoidea | ✓ | 14,479 | 382 | ✓ | 999 | 3,270 | GCA_000836215.1 |
| P. xuthus | Papilionoidea | ✓ | 12,723 | 384 | ✓ | 1,549 | 3,339 | GCA_000836235.1 |
| P. zelicaon | Papilionoidea | ✓ | 386 | TSA | ||||
| Spodoptera exigua | Noctuoidea | ✓ | 302 | TSA | ||||
| S. frugiperda | Noctuoidea | ✓ | 19,971 | 313 | ✓ | 1,350 | 3,315 | GCA_002213285.1 |
| S. littoralis | Noctuoidea | ✓ | 316 | TSA | ||||
| S. litura | Noctuoidea | ✓ | 30,212 | 317 | ✓ | 2,736 | 3,274 | GCA_002706865.1 |
| Vanessa tameamea | Papilionoidea | ✓ | 12 | 281 | ✓ | 5,456 | 3,364 | GCF_002938995.1 |
Overall, we got Persaeus/Zenon pair (i.e., the two elements from the same genome) from 12 species. On the other hand, for nine species we only got Persaeus and for six species we only found Zenon (these do not include H. melpomene and Bombyx mori for which the LINE was already known): Although in most cases this could be related to the databases where the species have been assayed, that could be limited and containing only repeat fragments such as nt, EST, or TSA databases, the exclusive presence of Persaeus or Zenon has been observed also in complete genomes (table 1).
Structure and Phylogenetic Analysis of Persaeus Elements
We collected a sample of 5, full-length copies of the Persaeus element from the 21 lepidopteran species in order to compare the sequence structure and variability. The resulting alignment can be partitioned in three main blocks: The 5′ and 3′ Zenon homologous regions and a variable central region (fig. 1). The two Zenon homologous regions showed a similar average nt identity of 66.0% and 67.6%, respectively. Moreover, a visual inspection of the alignment revealed a remarkable structural diversity among species, whereas repeats from congeneric species showed a more consistent structural pattern (fig. 1). The central variable region was found containing nt fragments that appear taxon-specific and whose homology among taxa do not seem obvious (fig. 1).
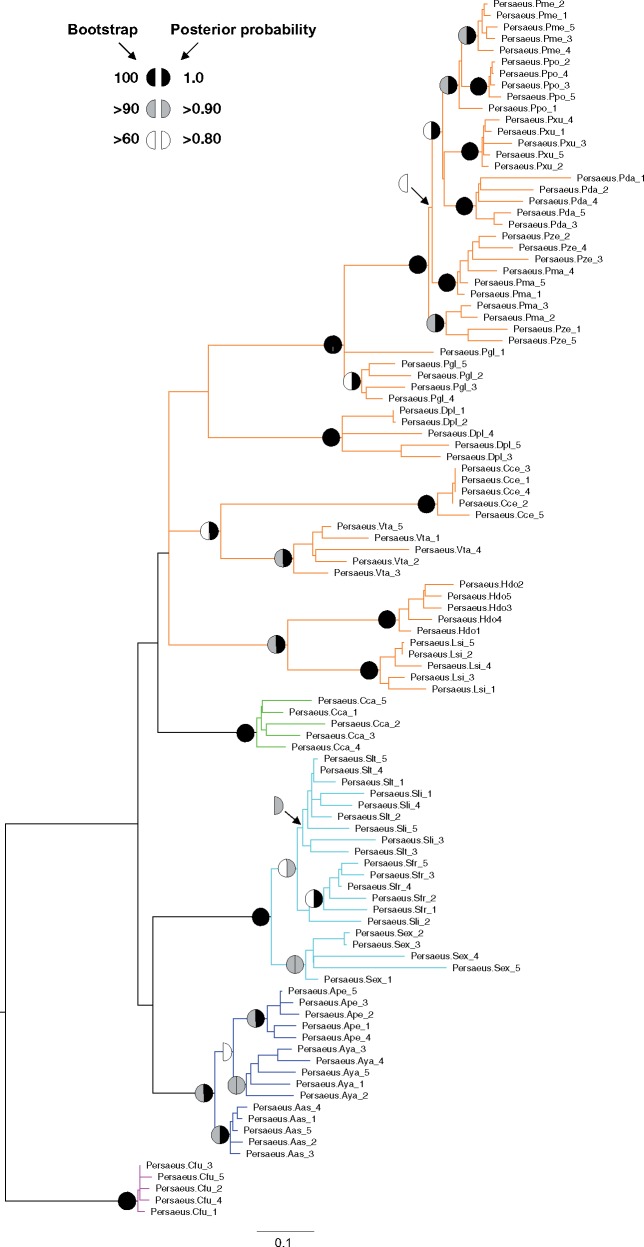
Maximum likelihood and Bayesian inference phylogenetic trees obtained using the 105 Persaeus copies resulted in two completely overlapping topologies: These are mostly unresolved at deep nodes but show higher support at the most recent nodes (fig. 2). Overall, SIDE sequences form species-specific clusters with the exception of repeats from Papilio machaon/P. zelicaon and Spodoptera litura/S. littoralis species pairs that are intermingled within their respective cluster (fig. 2). At genus level, SIDEs from Antheraea spp., Papilio spp., and Spodoptera spp. are included in the three, clearly monophyletic clades. At higher taxonomic level, Papilionoidea (the only superfamily for which more than one genus is available) are included in a single cluster, although not supported by maximum likelihood bootstrap or Bayesian posterior probabilities. Here, beside Papilio spp., two other species pairs are included in supported monophyletic groups: C. cecrops/Vanessa tameamea and Heliconius dori/Leptidea sinapis (fig. 2).
Fig. 2.
—Phylogenetic analysis Persaeus elements (five copies per species). Symbols at nodes represent maximum likelihood bootstrap/Bayesian posterior probability node support, as reported in the upper left legend. Branch color codes are indicative of the lepidopteran superfamily, as follow: Orange, Papilionoidea; blue, Bombycoidea; cyan, Noctuoidea; green, Pyraloidea; magenta, Tortricoidea. Each element has been labelled by a suffix indicating the pertaining species: Aya, Antheraea yamamai; Ape, A. pernyi; Aas, A. assama; Cca, Cadra cautella; Cce, Calycopis cecrops; Cfu, Choristoneura fumiferana; Dpl, D. plexippus; Hdo, H. doris; Lsi, Leptidea sinapis; Pda, Papilio dardanus; Pgl, P. glaucus; Pma, P. machaon; Pme, P. memnon; Ppo, P. polytes; Pxu, P. xuthus; Pze, P. zelicaon; Sex, Spodoptera exigua; Sfr, S. frugiperda; Slt, S. littoralis; Sli, S. litura; Vta, Vanessa tameamea.
Phylogenetic Analysis of Zenon Elements
We obtained full-length Zenon elements from 18 lepidopteran species. The RT protein domain was then used for phylogenetic analysis of newly isolated elements together with Zenon obtained from RepBase Update, Zenon-1_Hmel, Zenon-2_Hmel, Zenon-3_Hmel from H. melpomene and Zenon_BM from B. mori, and closely related CR1 elements from H. melpomene genome. Both maximum likelihood and Bayesian inference were congruent and are presented in supplementary figure S4, Supplementary Material online. The Zenon clade appeared monophyletic, although weakly supported; all Zenon elements for which a Persaeus SIDE has been isolated fell in the same supported cluster but intermingling with other Zenon elements obtained from genomes lacking the SIDE. As observed for Persaeus phylogeny, there are no clear relationships at superfamily taxonomic level but elements from congeneric species are consistently clustered together. The only exceptions are B. mori and B. mandarina elements that are paraphyletic with the remaining Zenon elements. Moreover, Heliconius spp. and L. sinapis elements are assembled in a supported cluster (supplementary fig. S4, Supplementary Material online).
Structural and Evolutionary Relationship between Persaeus and Zenon
In all SIDE/LINE pairs it is well clear the homology at the 5′ and 3′ end regions (supplementary dataset S1, Supplementary Material online). The nt identity between 5′ ends of each pair ranges from 71.0% in Papilio glaucus to 98.9% in V. tameamea, whereas the identity between 3′ ends ranges from 60.4% in Papilio polytes to 99.2% in V. tameamea (supplementary table S2, Supplementary Material online). In the Heliconius genomes, we only got Persaeus from H. doris, where Zenon was not observed; on the other hand, Zenon was found in the congeneric H. melpomene and H. numata. Despite they are present in different genomes, the identity between the homologous regions spans from 94.2% (Persaeus H. doris vs. Zenon-1_Hmel 3′ end) to 96.5% (Persaeus H. doris vs. Zenon-1_Hmel 5′ end) (supplementary table S2, Supplementary Material online). This holds also for Danaus spp. genomes, where Persaeus was found in D. plexippus but not D. chrysippus and vice versa for Zenon (table 1). Though, in this case, the identity at 5′ and 3′ ends dropped to 71.9% and 63.5%, respectively (supplementary table S2, Supplementary Material online).
The central variable region observed in Persaeus elements has no obvious similarity with respective LINEs, the nt fragments being scattered across the length of Zenon ORF with only small stretch of local similarity (supplementary dataset S1, Supplementary Material online).
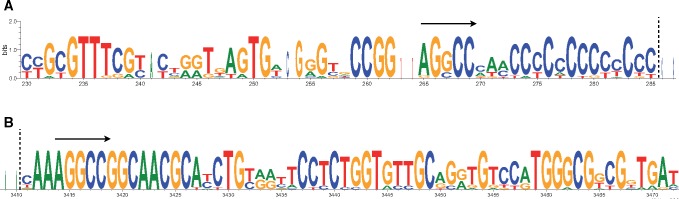
The 5′ end homologous region between Zenon and Persaeus terminates with a poly-(C) stretch (fig. 3) and it appears variable at break point among different SIDEs (fig. 1; supplementary dataset S1, Supplementary Material online); moreover, the Zenon’s region where internal deletion occurs is surrounded by 5 bp direct repeat 5′-AGGCC-3′ (fig. 3).
Fig. 3.
—Sequence logo of Zenon region surrounding the break point where internal deletions occur (dashed, vertical lines). (A) The region upstream the break point. (B) The region downstream the break point. Black arrows in (A) and in (B) indicate direct repeats (microhomologies).
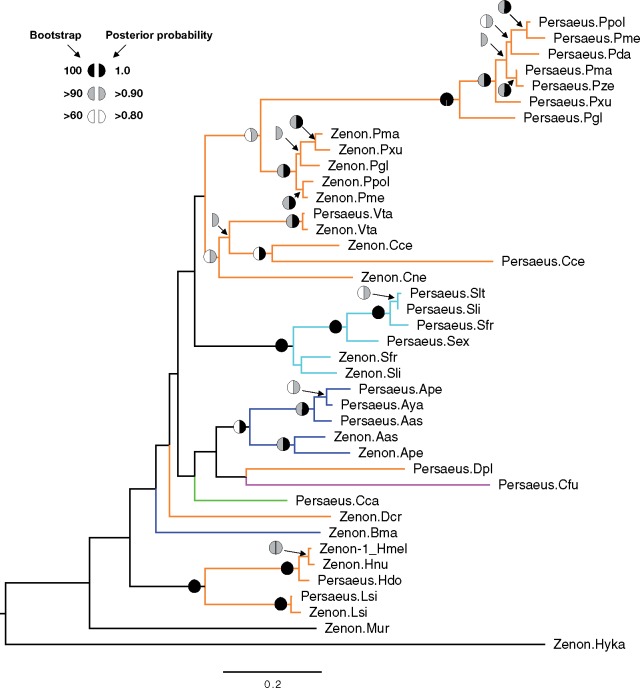
In order to determine the evolutionary relationship between Persaeus and Zenon, phylogenetic analyses were carried out based on Persaeus and Zenon consensus sequences (supplementary dataset S1, Supplementary Material online) using maximum likelihood and Bayesian inference. The two phylogenetic analyses are fully congruent and indicated a clustering pattern of Persaeus and Zenon not based on the host Lepidoptera superfamilies but based on host genus or species (Antheraea, Calycopis, Heliconius, Leptidea, Papilio, Spodoptera, and Vanessa). Only Persaeus and Zenon from Danaus spp. resulted more distantly related (fig. 4). Zenon and Persaeus from the genera represented by more than one species (Antheraea, Heliconius, Papilio, and Sopodoptera) not only cluster in monophyletic clades but each of these clades shows two further subclades, one for Zenon and one for Persaeus. Notably, the Persaeus subclades showed a topology that appears generally congruent with the species phylogeny (supplementary fig. S5, Supplementary Material online).
Fig. 4.
—Phylogenetic analyses of Persaeus and Zenon elements. Symbols at nodes represent support based on maximum likelihood bootstrap/Bayesian posterior probability, as reported on the upper left legend. Each element has been labelled by a suffix indicating the pertaining species: Aya, Antheraea yamamai; Ape, A. pernyi; Aas, A. assama; Bma, Bombyx mandarina; Cca, Cadra cautella; Cce, Calycopis cecrops; Cne, Calephelis nemesis; Cfu, Choristoneura fumiferana; Dcr, Danaus chrysippus; Dpl, D. plexippus; Hnu, Heliconius numata; Hdo, H. doris; Lsi, Leptidea sinapis; Hka, Hyposmocoma kahamanoa; Mur, Megathymus ursus; Pda, Papilio dardanus; Pgl, P. glaucus; Pma, P. machaon; Pme, P. memnon; Ppo, P. polytes; Pxu, P. xuthus; Pze, P. zelicaon; Sex, Spodoptera exigua; Sfr, S. frugiperda; Slt, S. littoralis; Sli, S. litura; Vta, Vanessa tameamea. Branch color codes as in figure 2.
When looking to activity through age analysis of Persaeus and Zenon in the same genome, we found an increase of Persaeus activity corresponding to the increased Zenon activity (fig. 5).
Fig. 5.
—Persaeus and Zenon activity through age analysis. Where necessary, data were magnified (as indicated in graph insets) in order to improve the readability.
Discussion
In this study, we identified a novel nonautonomous retroelement, the SIDE Persaeus, and analyzed the evolutionary dynamics with its partner CR1 LINE, Zenon, in Lepidoptera genomes. We also confirmed that CR1 retrotransposons, which are considered among the most abundant superfamily of TEs in the amniote genomes, are also abundant in insects, at least in Lepidoptera. Previous analyses already identified elements of the CR1 clade in insects, including Lepidoptera (Biedler and Tu 2003; Kapitonov and Jurka 2003; Novikova et al. 2007; Thompson et al. 2009). In the present analysis, we characterize the full-length sequence of additional 18 Zenon CR1 elements in further lepidopteran species. Most of LINEs, and especially CR1 elements, are frequently truncated at the 5′ end (Hillier et al. 2004), which make difficult to reconstruct the full-length CR1 as well as determine the exact boundary of their 5′ end. This, in part, explains why in some genomes we cannot retrieve full-length Zenon elements.
The finding and the evolutionary dynamics of the retrieved SIDE, Persaeus, are remarkable because this is, to our knowledge, the first instance of several independent successful genome invasions by an SIDE. Other SIDEs, such as R2 SIDE and R2/R1 hybrid SIDEs (Eickbush and Eickbush 2012), Ag-Sponge (Biedler and Tu 2003), and TbRIME (Kimmel et al. 1987) had been reported, but almost all these SIDEs as well as partner LINEs had low copy number in their host genomes. This has been attributed to various factors, among which the ability of the SIDE to be transcribed into RNA (Eickbush and Eickbush 2012). The other known successful nonautonomous retroelements, SINEs, are transcribed by the RNA pol-III thanks to the presence of promoter sequences in the RNA-related head (Luchetti and Mantovani 2013). The sequence of Persaeus 5′ end includes part of the CR1 5′-UTR, showing between 71% and 98% of identity, which is important for transcription because, in LINEs, it contains the promoter sequence (Lee et al. 2012). Therefore, Persaeus could retain the potential to be transcribed by the same mechanism of its partner Zenon. It has been showed that the 3′-UTRs of LINEs, including CR1 elements, is used as a recognition site for the encoded RT (Kajikawa et al. 1997; Haas et al. 2001; Suh 2015). Like the functional relationship between partner SINEs and LINEs, that is mediated by the similar nt sequence at the 3′ end (Ohshima and Okada 2005), Persaeus exhibited a 3′ end sharing 72–99% of identity with the Zenon 3′-UTR: Therefore, this suggests that it might borrow the retrotransposition machinery from its autonomous partner Zenon. This is also supported by the activity through age analysis, where Persaeus activity resulted contemporary to that of the partner Zenon in all assayed genomes.
Overall, at variance of previously identified SIDEs, it appears that Persaeus underwent to a replicative burst during Lepidoptera evolution reaching, on average, the 1.48% in length of the host genomes, with the remarkable instance of C. cecrops whose genome is made by the 3.48% of Persaeus SIDEs. Zenon activity showed the same trend, reaching an average genome coverage of 0.75% and with the maximum value scored in L. sinapis (3.45%).
Although the general structure of Persaeus is conserved across species, the sequences comparison revealed a more complex pattern. First of all, the regions homologous to Zenon 5′ and 3′ ends showed different structures that are consistent among closely related species (e.g., like the congeneric ones) but well differentiated between distantly related taxa. Moreover, the alignment pattern of the central variable region, which do not show any clear relationships with Zenon or any other sequences, suggest a nonhomologous origin. The phylogenetic analysis performed on SIDEs and LINEs indicated a concordant pattern of evolution. In fact, Persaeus and Zenon elements isolated from species of the same genus always cluster together, forming an SIDE and an LINE subclade in each genus or species clade (Antheraea, Calycopis, Heliconius, Lepitdea, Papilio, Spodoptera, and Vanessa). Altogether, the nonhomologous sequence structure and the phylogenetic pattern suggests that, although widespread among lepidopteran, the emergence of Persaeus occurred multiple time by internal deletion of a clade-specific Zenon element. The alternative hypothesis of a single origin of Persaeus appears unlikely as, in that case, we would have observed in the phylogenetic tree a single, ancient split between Zenon and Perseus sequences and then their diversification in the different clades. This is, actually, exactly the pattern that can be observed within those clades where multiple congeneric species are present (i.e., Antheraea, Heliconius, Papilio, and Spodoptera; fig. 4), suggesting that Persaeus emerged by Zenon internal deletion early during the evolution of these clades and that, because then, the two elements diverged independently. Moreover, when looking at the branching pattern within the Antheraea, Papilio, and Spodoptera clades it appeared that the Persaeus phylogeny resulted generally similar that of the host species. Although the taxon sampling is not exhaustive, as it is limited to genomic/transcriptomic data available for these genera in the database, this would suggest that the SIDE emerged in the common ancestor of each genus and then it was inherited following a vertical pattern. TEs are able to be transmitted by horizontal transfer, although with different rates based on specific biological feature of the element itself and of the host organism (Scavariello et al. 2017). Recent surveys on insect TEs indicated a global high frequency of horizontal transfers, evidencing a particular tendency of Lepidoptera to be involved in these events (Peccoud et al. 2017; Reiss et al. 2019). Moreover, it was found that these events preferentially took place among closely related species (Peccoud et al. 2017). However, horizontal transfers of nonautonomous retrotransposons have been only rarely reported (Hamada et al. 1997; Piskurek and Okada 2007; Luchetti et al. 2016; Luchetti and Mantovani 2016): This is probably because the lack of the specific partner autonomous element in the landing genome do not allow the replication of the transferred element. In the case of Persaeus, though, the co-occurrence of similar active LINEs could make possible such hypothetical successful horizontal transfer. Our data, apparently, seem to rule out this possibility in the assayed genomes but the Persaeus/Zenon partnership identified in this study might also provide an ideal system to investigate these interactions.
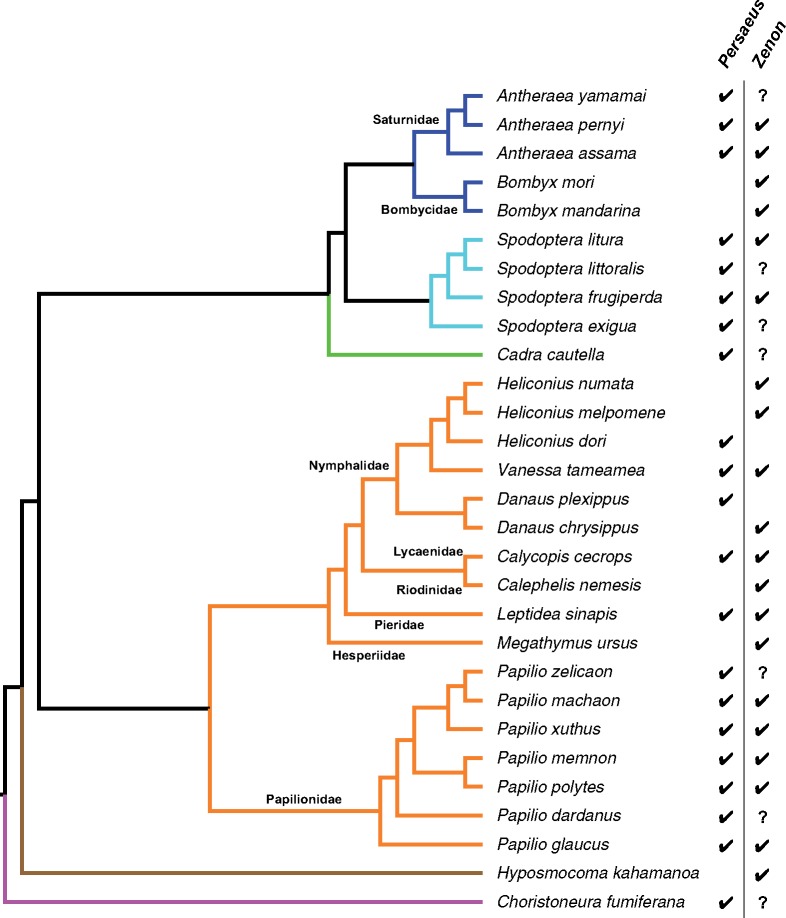
Although tested on a potentially limited taxon sampling, looking at the differential distribution of Persaeus and Zenon on the phylogeny of presently analyzed species (fig. 6), it appears that in some lineages the SIDE did not emerged or do not raised to a detectable copy number. However, the fact that Persaeus originated multiple times, even if with slightly different structure and from different member of the CR1 Zenon subfamily, indicates the presence some structural motif that may facilitate the internal deletion. Zenon sequence inspection evidenced the presence of short direct repeats bordering the region where internal deletion occurred. The presence of short direct repeats, also called microhomologies, has been thought to promote internal deletions among class II TE, through a DNA repair mechanism triggered after element excision (Rubin and Levy, 1997; Negoua et al., 2013). However, Zenon is a class I element where excisions, although possible, are rare events (van de Lagemaat et al. 2005). Another possible explanation for the frequent emergence of internal deletion derivatives could rely on recombination: Microhomologies could serve as nonhomologous sequences pairing region and recombination may occur. Interestingly, this could happen also during the reverse transcription process, as described in the copy-choice RNA recombination model: The RT enzyme is able to switch RNA template (template jump) between region of sequence similarity, leading to chimeric molecules (Simon-Loriere and Holmes 2011). This model has been repeatedly reported as potential generator of new SINE elements (Szafranski et al. 2004; Luchetti and Mantovani 2016). Moreover, the RT enzyme could be able to add nontemplate nt while template jumping (Bibillo and Eickbush 2004): This could possibly explain the presence of nt stretches in the central variable region of Persaeus which are not clearly related to other elements’ sequences.
Fig. 6.
—Summary of Persaeus and Zenon distribution across the hosts phylogeny. Question marks indicate species where genome sequence was not available and, therefore, the absence of Zenon maybe to the limited data set available (GenBank database EST, TSA, or nt; table 1).
Overall, the evolutionary consequences of the amplification burst of Persaeus and Zenon found here need to be further investigated, even because the invasion of substantial fraction of DNA generated by transposition of TEs can strongly affect the structure and functionality of genomes (Feschotte and Pritham 2007; Cordaux and Batzer 2009). CR1 transposons are widespread in the genomes of amniotes and they were the only active transposons during the avian lineage evolution (Hillier et al. 2004; Kaiser et al. 2006; Warren et al. 2010; Haddrath and Baker 2012; Liu et al. 2012; Baker et al. 2014). The characteristics of widespread distribution and high copy number of Persaeus and Zenon seem to imply that CR1 elements are also active throughout Lepidoptera evolution.
Supplementary Materials
Supplementary data are available at Genome Biology and Evolution online.
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Ethics Approval
Not applicable.
Funding
This work was supported by the Funds for Distinguished Young Scientists of Jiangxi Province (20192BCBL23028), the National Natural Science Foundation of China (31700318 and 31560308) to H.H.Z and by Canziani funding to A.L.
Authors’ Contributions
H.H.Z., P.L.W., and A.L. designed and supervised the study. A.L., A.A.R., P.L.W., M.R.X.X., X.M.X., X.G.Z., and H.H.Z. performed bioinformatic analyses. P.L.W., X.M.X., X.G.Z., H.H.Z., and A.L. wrote and revised the manuscript.
Supplementary Material
Literature Cited
- Bailly-Bechet M, Haudry A, Lerat E.. 2014. One code to find them all: a Perl tool to conveniently parse RepeatMasker output files. Mob DNA 5(1):13. [Google Scholar]
- Baker AJ, Haddrath O, McPherson JD, Cloutier A.. 2014. Genomic support for a moa-tinamou clade and adaptive morphological convergence in flightless ratites. Mol Biol Evol. 31(7):1686–1696. [DOI] [PubMed] [Google Scholar]
- Bibillo A, Eickbush T.. 2004. End-to-end template jumping by the reverse transcriptase encoded by the R2 retrotransposon. J Biol Chem. 279(15):14945–14953. [DOI] [PubMed] [Google Scholar]
- Biedler J, Tu Z.. 2003. Non-LTR retrotransposons in the African malaria mosquito, Anopheles gambiae: unprecedented diversity and evidence of recent activity. Mol Biol Evol. 20(11):1811–1825. [DOI] [PubMed] [Google Scholar]
- Burch JBE, Davis DL, Haas NB.. 1993. Chicken repeat 1 elements contain a pol-like open reading frame and belong to the non-long terminal repeat class of retrotransposons. Proc Natl Acad Sci U S A. 90(17):8199–8203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castoe TA. 2013. The Burmese python genome reveals the molecular basis for extreme adaptation in snakes. Proc Natl Acad Sci U S A. 110(51):20645–20650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cordaux R, Batzer MA.. 2009. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 10(10):691–703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eickbush DG, Eickbush TH.. 2012. R2 and R2/R1 hybrid non-autonomous retrotransposons derived by internal deletions of full-length elements. Mob DNA 3(1):10.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feschotte C, Pritham EJ.. 2007. DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet. 41:331–368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao D, Li Y, Kim KD, Abernathy B, Jackson SA.. 2016. Landscape and evolutionary dynamics of terminal repeat retrotransposons in miniature in plant genomes. Genome Biol. 17:7.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green RE, et al. 2014. Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346(6215):1254449.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haas NB, et al. 2001. Subfamilies of CR1 non-LTR retrotransposons have different 5′UTR sequences but are otherwise conserved. Gene 265(1–2):175–183. [DOI] [PubMed] [Google Scholar]
- Haas NB, Grabowski JM, Sivitz AB, Burch J.. 1997. Chicken repeat 1 (CR1) elements, which define an ancient family of vertebrate non-LTR retrotransposons, contain two closely spaced open reading frames. Gene 197(1–2):305–309. [DOI] [PubMed] [Google Scholar]
- Haddrath O, Baker AJ.. 2012. Multiple nuclear genes and retroposons support vicariance and dispersal of the palaeognaths, and an early cretaceous origin of modern birds. Proc Biol Sci. 279(1747):4617–4625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamada M, et al. 1997. A newly isolated family of short interspersed repetitive elements (SINEs) in coregonid fishes (whitefish) with sequences that are almost identical to those of the SmaI family of repeats: possible evidence for the horizontal transfer of SINEs. Genetics 146(1):355–367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hillier LW, et al. 2004. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432:695–716. [DOI] [PubMed] [Google Scholar]
- Jurka J. 2000. Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16(9):418–420. [DOI] [PubMed] [Google Scholar]
- Kaiser VB, van Tuinen M, Ellegren H.. 2006. Insertion events of CR1retrotransposable elements elucidate the phylogenetic branching order in galliform birds. Mol Biol Evol. 24(1):338–347. [DOI] [PubMed] [Google Scholar]
- Kajikawa M, Ohshima K, Okada N.. 1997. Determination of the entire sequence of turtle CR1: the first open reading frame of the turtle CR1 element encodes a protein with a novel zinc finger motif. Mol Biol Evol. 14(12):1206–1217. [DOI] [PubMed] [Google Scholar]
- Kapitonov VV, Jurka J.. 2003. The esterase and PHD domains in CR1-Like non-LTR retrotransposons. Mol Biol Evol. 20(1):38–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J, Mun S, Meyer TJ, Han K.. 2012. High levels of sequence diversity in the 5′ UTRs of human-specific L1 elements. Comp Funct Genomics. 2012:129416.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luchetti A, Mantovani B.. 2013. Conserved domains and SINE diversity during animal evolution. Genomics 102(4):296–300. [DOI] [PubMed] [Google Scholar]
- Luchetti A, Mantovani B.. 2016. Rare horizontal transmission does not hide long-term inheritance of SINE highly conserved domains in the metazoan evolution. Curr Zool. 62(6):667–674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luchetti A, Satovic E, Mantovani B, Plohl M.. 2016. RUDI, a short interspersed element of the V-SINE superfamily widespread in molluscan genomes. Mol Genet Genomics. 291(3):1419–1429. [DOI] [PubMed] [Google Scholar]
- Kergoat GJ, et al. 2012. Disentangling dispersal, vicariance and adaptive radiation patterns: a case study using armyworms in the pest genus Spodoptera (Lepidoptera: noctuidae). Mol Phylogenet Evol. 65(3):855–870. [DOI] [PubMed] [Google Scholar]
- Kimmel BE, Ole-Moiyoi OK, Young JR.. 1987. Ingi, a 5.2-kb dispersed sequence element from Trypanosoma brucei that carries half of a smaller mobile element at either end and has homology with mammalian LINEs. Mol Cell Biol. 7(4):1465–1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Z, He L, Yuan H, Yue B, Li J.. 2012. CR1 retroposons provide a new insight into the phylogeny of Phasianidae species (Aves: galliformes). Gene 502(2):125–132. [DOI] [PubMed] [Google Scholar]
- Negoua AH, Rouault J-D, Chakir M, Capy P.. 2013. Internal deletions of transposable elements: the case of Lemi elements. Genetica 141(7–9):369–379. [DOI] [PubMed] [Google Scholar]
- Novikova O, et al. 2007. CR1 clade of non-LTR retrotransposons from Maculinea butterflies (Lepidoptera: lycaenidae): evidence for recent horizontal transmission. BMC Evol Biol. 7:93.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohshima K, Okada N.. 2005. SINEs and LINEs: symbionts of eukaryotic genomes with a common tail. Cytogenet Genome Res. 110(1–4):475–490. [DOI] [PubMed] [Google Scholar]
- Peccoud J, Loiseau V, Cordaux R, Gilbert C. 2017. Massive horizontal transfer of transposable elements in insects. Proc Natl Acad Sci U S A.114(18):4721–4726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piskurek O, Okada N.. 2007. Poxviruses as possible vectors for horizontal transfer of retroposons from reptiles to mammals. Proc Natl Acad Sci U S A. 104(29):12046–12051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reiss D, et al. 2019. Global survey of mobile DNA horizontal transfer in arthropods reveals Lepidoptera as a prime hotspot. PLoS Genet. 15(2):e1007965.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ronquist F, et al. 2012. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 61(3):539–542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin E, Levy AA.. 1997. Abortive gap repair: underlying mechanism for Ds element formation. Mol Cell Biol. 17(11):6294–6302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scavariello C, Luchetti A, Bonandin L, Martoni F, Mantovani B.. 2017. Hybridogenesis and a potential case of R2 non-LTR retrotransposon horizontal transmission in Bacillus stick insects (Insecta Phasmida). Sci Rep. 7(1):41946.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaffer HB, et al. 2013. The western painted turtle genome, a model for the evolution of extreme physiological adaptations in a slowly evolving lineage. Genome Biol. 14(3):R28.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simon-Loriere E, Holmes EC.. 2011. Why do RNA viruses recombine? Nat Rev Microbiol. 9(8):617–626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh D, et al. 2017. The mitochondrial genome of Muga silkworm (Antheraea assamensis) and its comparative analysis with other lepidopteran insects. PLoS One 12(11):e0188077.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smit AFA, Hubley R, Green P.. 2013. –2015. RepeatMasker Open-4.0. Available from: http://www.repeatmasker.org.
- Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stumph WE, Hodgson CP, Tsai MJ, O'Malley BW.. 1984. Genomic structure and possible retroviral origin of the chicken CR1 repetitive DNA sequence family. Proc Natl Acad Sci U S A. 81(21):6667–6671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stumph WE, Kristo P, Tsai M-J, O'Malley BW.. 1981. A chicken middle-repetitive DNA sequence which shares homology with mammalian ubiquitous repeats. Nucleic Acids Res. 9(20):5383–5398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suh A. 2015. The Specific Requirements for CR1 retrotransposition explain the scarcity of retrogenes in birds. J Mol Evol. 81(1–2):18–20. [DOI] [PubMed] [Google Scholar]
- Suh A, et al. 2014. Multiple lineages of ancient CR1 retroposons shaped the early genome evolution of amniotes. Genome Biol Evol. 7(1):205–217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szafranski K, Dingermann T, Glockner G, Winckler T.. 2004. Template jumping by a LINE reverse transcriptase has created a SINE-like 5S rRNA retropseudogene in Dictyostelium. Mol Genet Genomics. 271(1):98–102. [DOI] [PubMed] [Google Scholar]
- Thompson ML, Gauna AE, Williams ML, Ray DA.. 2009. Multiple chicken repeat 1 lineages in the genomes of oestroid flies. Gene 448(1):40–45. [DOI] [PubMed] [Google Scholar]
- van de Lagemaat LN, Gagnier L, Medstrand P, Mager DL.. 2005. Genomic deletions and precise removal of transposable elements mediated by short identical DNA segments in primates. Genome Res. 15(9):1243–1249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warren WC, et al. 2010. The genome of a songbird. Nature 464(7289):757–762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zakharov EV, Caterino MS, Sperling F.. 2004. Molecular phylogeny, historical biogeography, and divergence time estimates for swallowtail butterflies of the genus Papilio (Lepidoptera: papilionidae). Syst Biol. 53(2):193–215. [DOI] [PubMed] [Google Scholar]
- Zhang HH, Feschotte C, Han MJ, Zhang Z.. 2014. Recurrent horizontal transfers of Chapaev transposons in diverse invertebrate and vertebrate animals. Genome Biol Evol. 6(6):1375–1386. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.