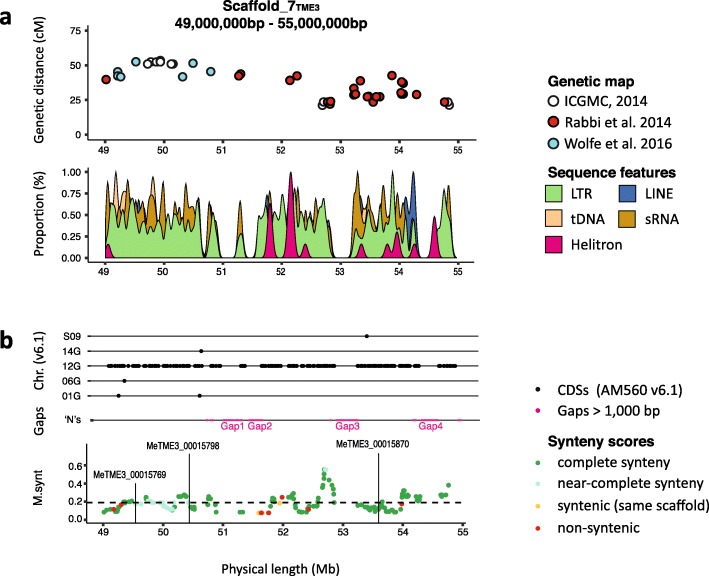
Fig. 5.
CMD2 locus in TME3 genome. a The upper panel shows CMD2-associated genetic SNP markers and their genetic distance relative to their physical position on scaffold_7 of TME3. Red dots indicate CMD2 SNP markers released by Rabbi and colleagues [16], and blue dots indicate the SNP markers released by Wolfe and colleagues [22, 42]. The lower panel shows the distribution of main repetitive genomic features at the CMD2 locus. b The upper panel shows the alignment position of AM560 v6.1 CDS in the region of Chr. 12 containing the CMD2 locus. Each black dot represents the CDS alignment position at the CMD2 scaffold (x-axis) and its chromosomal origin from the AM560 v6.1 cassava reference genome. Sequence breaks (gaps > 1 Kb) are shown as pink bars. The lower panel shows the MSS for every annotated gene at the CMD2 locus in TME3. Green dots indicate genes that are found in the CMD2 region of 60444, and light blue dots indicate genes that are found in close proximity of the CMD2 locus in 60444. Orange dots indicate TME3 genes that show a syntenic relation to 60444 genes on other 60444 scaffolds, and red dots indicate genes with no syntenic relation. The dashed line represents the MSS average for the whole genome