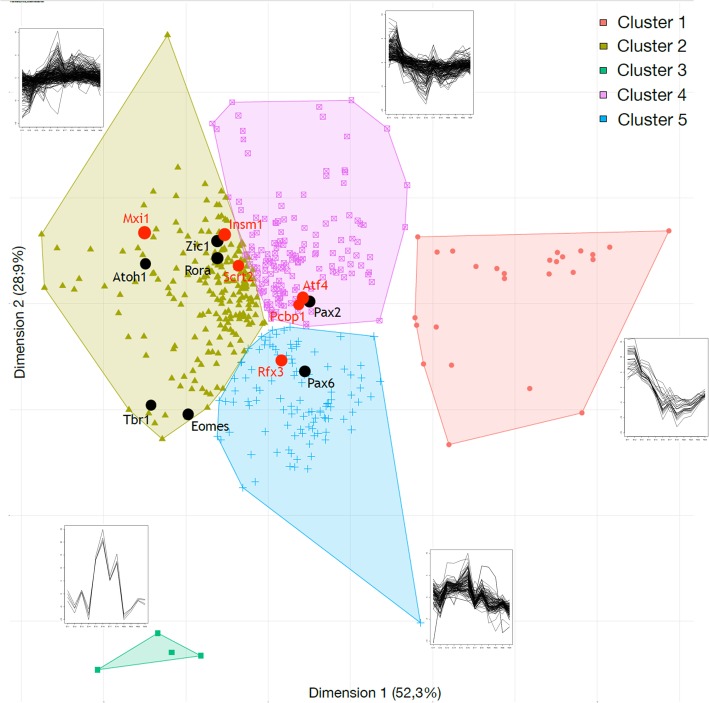
Fig. 3.
K-mean clustering of TFs based on their motif activity pattern in cerebellar development. Transcription factors are clustered based on their temporal motif activity pattern in cerebellar development. The motif activity profiles are shown beside each cluster. Genes highlighted with black dots have known functional roles during cerebellar development. Potential novel regulators with similar motif activity patterns can be identified as a neighbor to known genes. Six candidates – Atf4, Insm1, Mxi1, Pcbp1, Rfx3 and Scrt2 that were used for validation experiments, highlighted with red dots, are neighbors to known cerebellar regulators - Atoh1, Pax2, Pax6, Rora and Zic1, highlighted with black dots