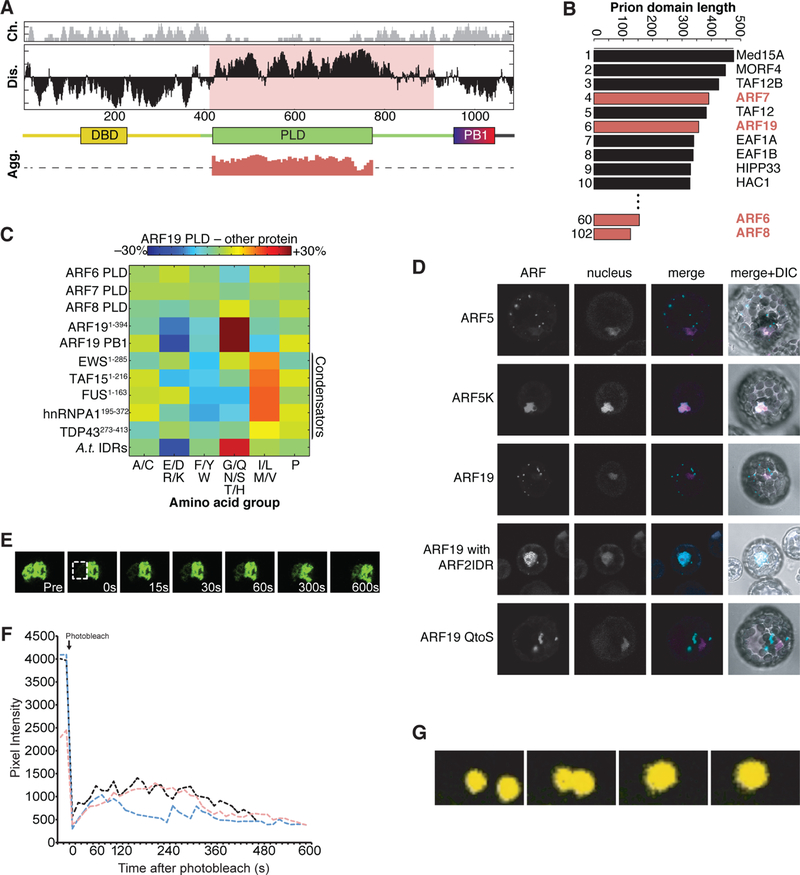
Figure 3. Sequence features of ARF19 and relationship to other condensate-forming proteins.
(A) Integrative linear sequence analysis showing local charge density (Ch.), local disorder propensity (Dis.) domain structure, and simulation-derived aggregation propensity of prion-like domain (PLD) alone (Agg.). The domain structure highlights the two known functional domains (DBD and PB1) and the PLD. The MR is predicted to be disordered and contains a PLD that has a strong propensity to self-associate.
(B) Proteome-wide analysis shows the top ten PLDs by length in the A. thaliana non-redundant proteome. See Dataset S1 for full list of Arabidopsis PRD-containing proteins.
(C) Comparative analysis of the ARF19 PLD composition in relation to different proteins. Each amino acid is assigned to one of six groups on the X-axis and the fractional difference of grouped amino acids between the ARF19 PLD and each sequence is shown. All sequence comparisons are in reference to the ARF19 PLD in relation to sequences from additional ARF PLDs, ARF19 DBD and PB1 domains, and PLD domains from known condensators. The average intrinsically disordered region (IDR) signature of disordered proteins in A. thaliana is also provided for reference.
(D) Confocal images of Wt (Col-0) protoplasts transfected with UBQ10:mVenus-ARF5, UBQ10:mVenus-ARF5K797A, UBQ10:mVenus-ARF19, UBQ10:mVenus-ARF19w/ARF2MR, and UBQ10:mVenus-ARF19QtoS. Co-transfection with a 35S:NLS-2*mCerulean marker was used to visualize nuclei.
(E) Time course of FRAP of condensate in UBQ10:YFP-ARF19 seedling. Dashed box indicates the initial photobleached region.
(F) Fluorescence recovery curve of three independent FRAP experiments on UBQ10:YFP-ARF19 condensates.
(G) Time course showing fusion of cytoplasmic condensates in root transition zone cells (containing both nuclear and condensate signal) in a seedling expressing ARF19:ARF19-Venus.