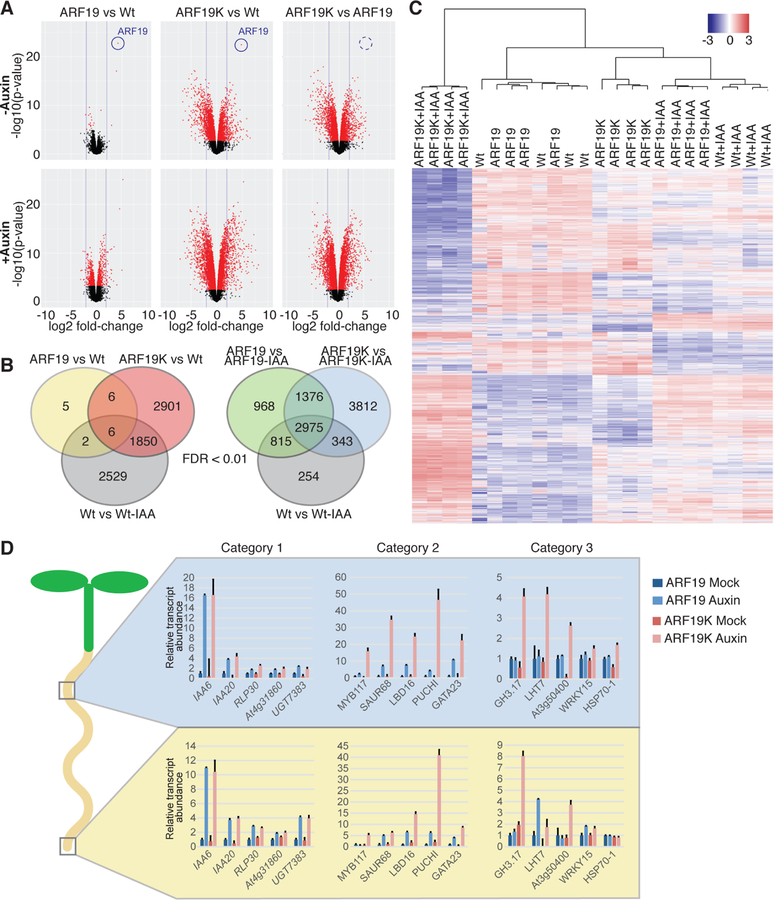
Figure 5. Disruption of ARF condensate formation results in an altered transcriptional landscape and morphological defects.
(A) Volcano plots displaying pairwise transcript accumulation differences between UBQ10:YFP-ARF19 (ARF19) and Wt (Col-0), between UBQ10:YFP-ARF19K962A (ARF19K) and Wt, and between ARF19K and ARF19 lines, each ± 10 µM IAA treatment. Transcripts whose accumulation differences are significant (FDR ≤ 0.01) are represented in red.
(B) Venn diagrams showing the overlap between the data sets of differentially expressed genes (FDR <0.01).
(C) Hierarchical clustering of genes displaying differential expression among samples (FDR <0.01).
(D) Relative transcript abundance (+SE; n = 3) of selected genes obtained by Nanostring analysis on upper root and lower root sections from 7d-old mock and auxin-treated UBQ10:YFP-ARF19 and UBQ10:YFP-ARF19K962A seedlings. Category 1 – auxin responsive in upper and lower root of ARF19 and ARF19K, Category 2 – auxin responsive in upper and lower root of ARF19 and ARF19K with a higher amplitude of change in auxin-treated ARF19K, Category 3 – auxin responsive only in the lower root of ARF19, but in both the upper and lower root of ARF19K + auxin responsive in only ARF19K.
See Figures S6 and S7 for RNASeq quality assessments. See Dataset S2 for pairwise comparisons of mock-treated samples, Dataset S3 for pairwise comparisons of auxin-treated samples, and Dataset S4 for comparisons between mock and auxin treatment within each line.