Figure 7. Disruption of mSWI/SNF Complex Assembly in Human Disease.
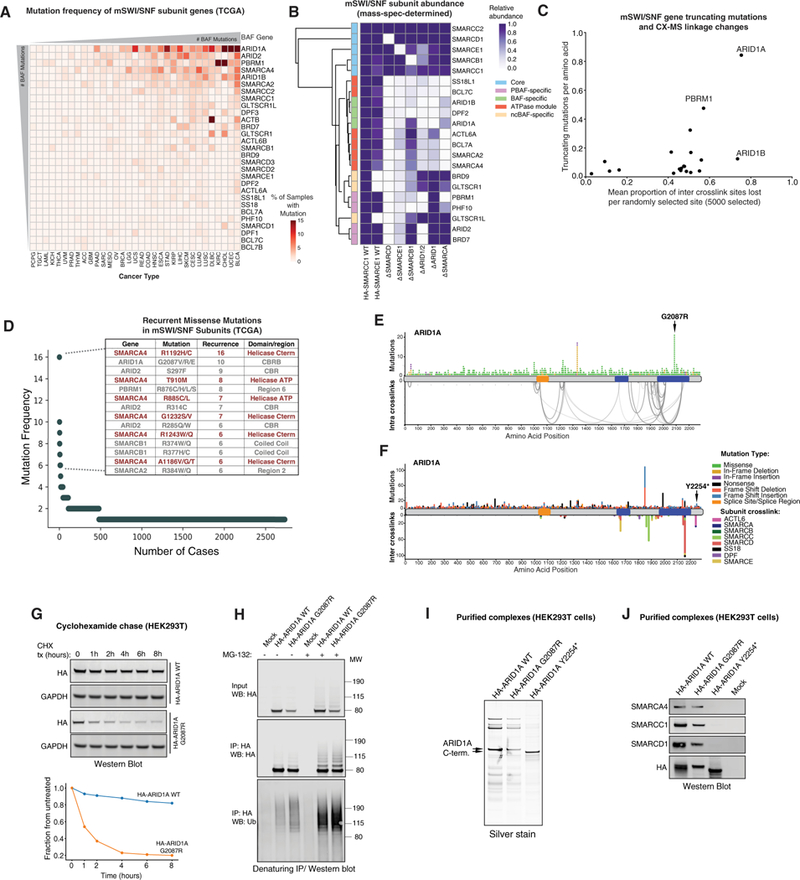
(A) Frequency of mSWI/SNF gene mutations across human cancers (The Cancer Genome Atlas [TCGA]). Hypermutated samples were excluded from analysis.
(B) Mass spectrometry analysis of mSWI/SNF complex subunit relative abundance in complexes purified from the indicated cell types (WT and subunit KO cells), normalized toWTSMARCC1 purifications. DSMARCDcomplexeswere purified using SMARCE1; ∆SMARCE1, ∆SMARCB1, ∆ARID1/2, ∆ARID1, and ∆SMARCA complexes were purified using HA-SMARCD1.
(C) Correlation analysis reflectingtheeffect oftruncating mutationson mSWI/SNF subunit linkages. Subunits mostfrequentlytruncated exhibit higherproportions of inter-crosslinked sites lost.
(D) Top-ranked cancer-associated missense mutations (TCGA). Mutations predicted to disrupt catalytic activity are shown in red.
(E) Non-truncating mutations in ARID1A across human cancers mapped over intra-crosslinks. The hotspot mutation in the highly crosslinked C-terminal CBRB region of the protein is indicated.
(F) Truncating mutations in ARIDIAacross human cancers mapped over crosslinks to other BAF subunits. The position ofthetruncating mutation Y2254* used in this study is indicated by the arrow.
(G) Top: Representative cycloheximide chase experiment assessing the half-life of ARID1A WT and G2087R mutant C-terminal region variants. Bottom: quantification of western blot (WB) above, normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH).
(H) MG-132 treatment (8 hr) of HEK293T cells expressing ARID1A WT and G2087R C-terminal regions. IP performed under denaturing conditions; immunoblot performed using indicated antibodies.
(I) Silver stain performed on ARID1A WT, G2087R, and Y2254* BAF complexes purified from HEK293T cells.
(J) Immunoblot of ARID1A WT, G2087R-, and Y2254*-bound BAF complexes purified from HEK293T cells.
See also Figure S7.
