Figure 2. SsoMCM orientation mapping onto 3’-(DNA171/165-3) or 5’-(DNA172/164-5) long arm fork DNA substrates.
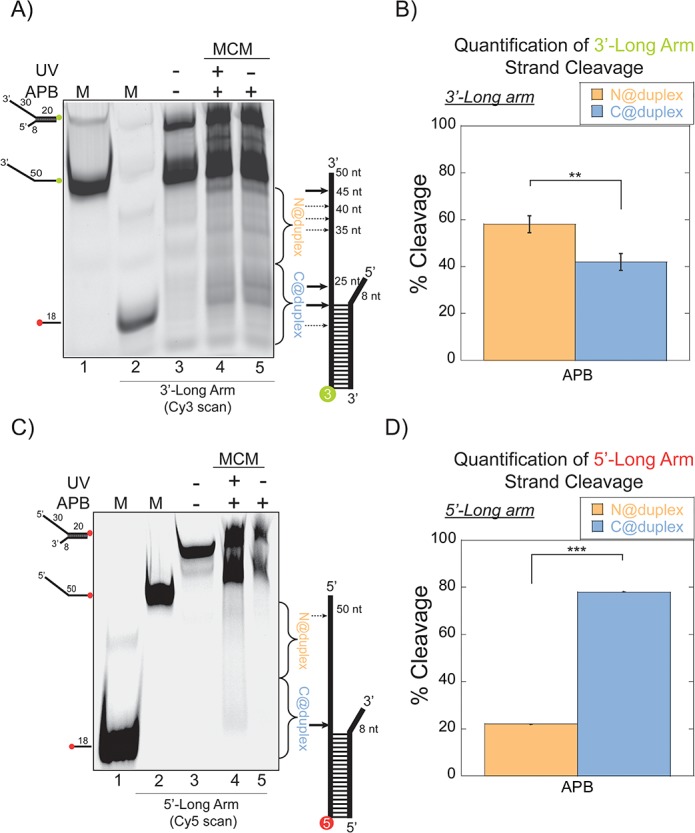
(A) APB orientation mapping of the 3’-encircled strand labelled at the 5’ duplex end with Cy3 on a 3’-long arm fork DNA substrate with a 20 base duplex. SsoMCM was labelled with APB at C682 (within the C-term WH motif) specifically. (B) Quantification of the relative amount of DNA cleavage for bases 20–35 or 36–50 from the 5’-end indicate the relative orientation for placing the N@duplex (orange) or C@duplex (blue), respectively closer to the duplex junction. Similarly, (C) APB orientation mapping of the 5’-excluded strand labelled at the 3’-duplex end with Cy5 on a 5’-long arm fork DNA substrate with a 20 base duplex. (D) Quantification of the relative amount of DNA cleavage for bases 20–35 or 36–50 from the 3’-end indicate the relative orientation for N@duplex or C@duplex, respectively, closer to the duplex junction. DNA cleavage was induced and enhanced with UV light [(A), lane 4, (C), lane 4]. Arrow thickness indicates the relative amount and position of DNA cleavage. DNA markers (M) indicate 18 and 50 bases and fork DNA. Error bars represent standard error from 3 to 5 independent experiments. The products were run on a 20% native PAGE gel. p-values are defined as *<0.05, **<0.01 ***<0.001. shorter 8 nt 3’-arm (Figure 2C–D). Here, the footprinting orientations were reversed, with a > 3:1 preference for C@duplex (Figure 2D). Therefore, on these long arm fork DNA substrates, SsoMCM can bind either the 3’- or 5’-arm in both orientations, but the preferred 3’−5’ polarity is CTD-NTD.
