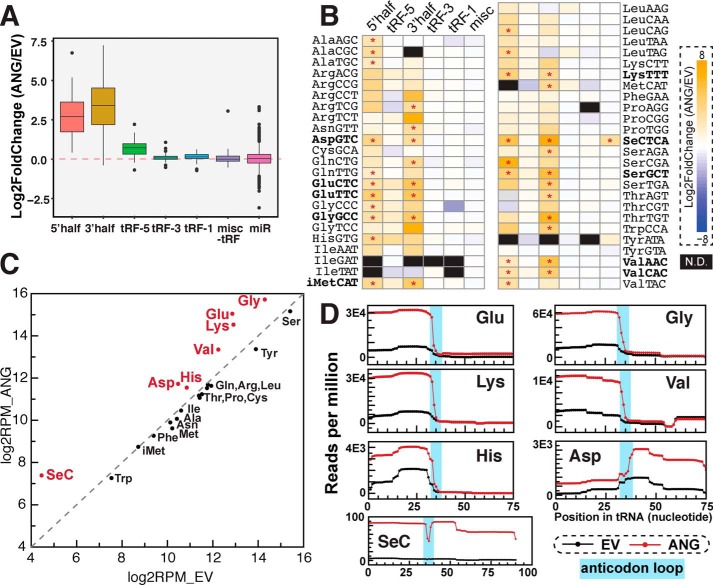
Figure 2.
tRNA fragments up-regulated by angiogenin overexpression. A–D, tRF changes upon ANG overexpression by small RNA-Seq. A and B, tRNA halves are increased by ANG overexpression. A, Box-and-whisker plot showing relative -fold change of reads per million mapped reads on log2 scale of ANG overexpression compared with the empty vector control (averaged from three biological replicates). tRFs are grouped by types and parental tRNAs. Box plot shows median value as middle line, first and third quartile as box outline, and data range (excluding outliers represented by individual dots) as whiskers. B, heat map shows log2 -fold change of tRFs by ANG overexpression. tRFs are grouped by tRF types and parental tRNAs. Significant changes determined by DESeq2 (adjusted p value less than 0.05) are labeled by asterisks. N.D., not detected. C and D, ANG overexpression increases tRF reads from certain parental tRNAs, including tRNAGlu, tRNAGly, tRNALys, tRNAVal, tRNAHis, tRNAAsp, and tRNASeC. For C, tRF reads (all types) were summed by parental tRNAs (x-axis: empty vector, y-axis: ANG overexpression). For D, coverage plot shows where accumulated tRF reads (excluding tRF-1s) map to the parental mature tRNAs (black line: empty vector control, red line: ANG overexpression). Anticodon loop region for each tRNA (predicted by tRNA covariance model -fold in GtRNAdb) is highlighted in cyan.