Fig. 3.
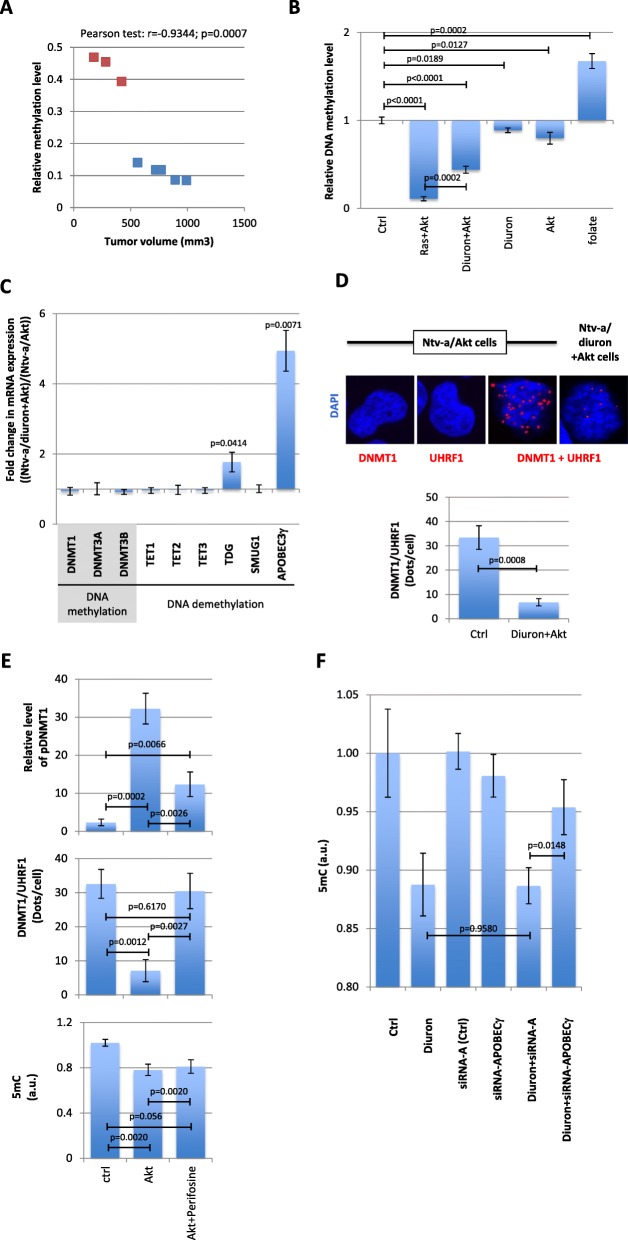
Global DNA hypomethylation occurs in Akt + diuron-induced glioma. a Correlation between the relative level of DNA methylation and the tumor volume. Blue squares represent the Ras + Akt-induced glioma (n = 5) and red squares represent the Akt + diuron-induced glioma (n = 3). b Graph illustrates the relative DNA methylation level (mean ± SD) seen in Ntv-a/Ras + Akt- and Ntv-a/diuron + Ak cells (n = 3). “Folate” represents the relative DNA methylation level in cells treated with folate, i.e., as a positive control of gain of DNA methylation. “Ctrl” represented the methylation level of Ntv-a/lacZ cells. c Graph represents the fold change expression in mRNA encoding for DNA methylation players (DNMT1, 3A, and 3B) and DNA demethylation players (TET1, 2, and 3, TDG, SMUG1, and APOBEC3γ). d ApoTome view of the DNMT1/UHRF1 dots in indicated cells. Red dots symbolize the DNMT1/UHRF1 interactions. Nucleus/DNA are stained in blue via the use of DAPI. Graph (mean ± SEM) illustrating the number of DNMT1/UHRF1 dots per cells in indicated cells. The number of DNMT1/UHRF1 dots is calculated from the analysis of, at least, 50 nucleus in three independent experimentations. The absence of dots in the presence of the unique use of DNMT1 or UHRF1 antibodies underlines the specificity of signal obtained in presence of the DNMT1 + UHRF1 antibodies. e Graphs illustrate the effect of Akt overexpression and/or inhibition (via the use of Perifosine) on the relative level of phospho-DNMT1 (pDNMT1) (top), the DNMT1/UHRF1 interactions (middle), and on the global level of 5-methylcytosine (5mC, bottom). P-LISA estimates the number of DNMT1/UHRF1 interactions. In cell ELISA performed with anti-pDNMT1S127 was used to analyze the DNMT1 phosphorylation such as previously described [3]. ELISA-5mC (Zymo Research, France) was used to quantify the 5methylcytosine (5mC), i.e., to study the global level of DNA methylation. f Graph illustrates the modification of 5-methylcystosine seen in cells treated with diuron and/or siRNA directed against APOBEC3γ. ELISA-5mC (Zymo Research, France) was used to quantify the 5methylcytosine (5mC), i.e., to study the global level of DNA methylation