Fig. 1.
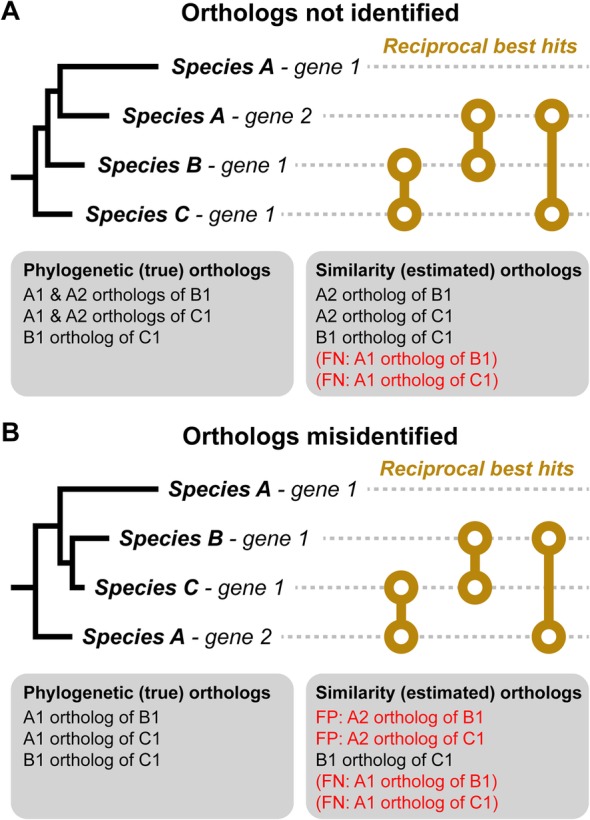
Pairwise similarity score-based ortholog inference can be misled by variable sequence evolution rates. a Phylogenetic tree of a typical gene family. The correct orthologs of species A-gene 1 are not identified. b Species A-gene 2 is misidentified as the ortholog of the genes from species B and C. Left-hand side: gene trees with branches to scale and the true orthology relationships, which can be determined from the gene tree. Right-hand side: Reciprocal best hits (RBH) based on gene similarity scores that are monotonic with branch length and the orthology relationships inferred from these scores using standard heuristics (orthologs inferred using RBHs and co-orthology identified from within species hits better than closest RBH [8, 16]). FP, false positive; FN, false negative
