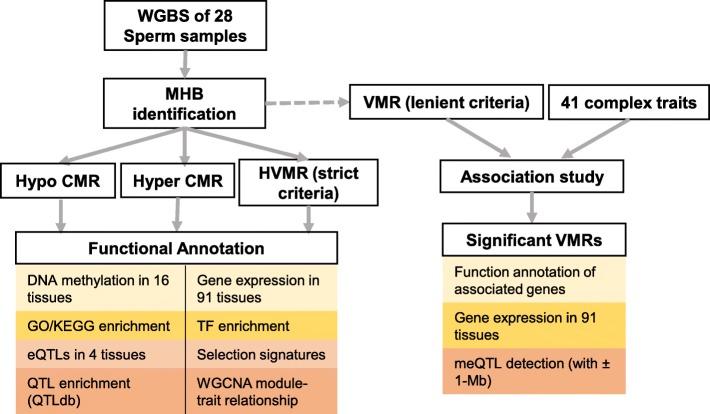
Fig. 1.
Schematic overview of the current study. We defined methylation haplotype blocks (MHBs) using whole genome bisulfite sequencing (WGBS) data of 28 sperm samples. We then detected the highly variably methylated regions (HVMRs), conserved hypomethylated regions (Hypo-CMRs) (average methylation level < 20%) and conserved hypermethylated regions (Hyper-CMRs) (average methylation level > 80%) based on the methylation variations among individuals. We next functionally annotated them by integrating DNA methylation, gene expression, GO/KEGG, transcriptional factor binding sites, QTL and WGCNA module-trait relationship. We further detected the variably methylated regions (VMRs) using lenient criteria. We associated the methylation levels of VMRs with 41 complex traits. We also annotated the significant VMRs by examining the functional annotation of their associated genes, and their corresponding expression across 91 tissues. We finally conducted cis-methylation QTL (± 1-Mb) analyses for significant VMRs