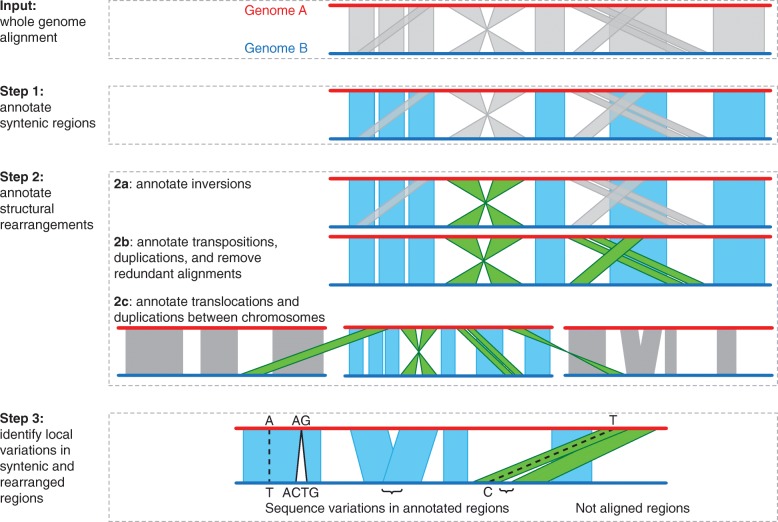
Fig. 2.
Workflow for the identification of genomic differences. SyRI uses whole-genome alignments (WGA) as input. A WGA consists of a set of local alignments, where each local alignment (gray polygon) connects a specific region in one genome to a specific region in the other genome. Step 1: SyRI identifies the highest scoring syntenic path between the corresponding genomes (blue alignments). The syntenic path represents the longest set of non-rearranged regions between two genomes. Step 2 (a–c): The remaining alignments are separated into structural rearrangements and redundant alignments. Structural rearrangements (green alignments) are classified into inversions, transpositions, and duplications, and finally inter-chromosomal rearrangements. Step 3: Local differences in the sequences are identified in all syntenic and rearranged regions. SNPs and small indels are parsed directly from the local alignments, whereas more complex sequence variations (e.g., like large indels and CNVs) are identified in the overlaps and gaps between consecutive local alignments. Also, all non-aligned regions in between syntenic and rearranged regions are reported for completeness