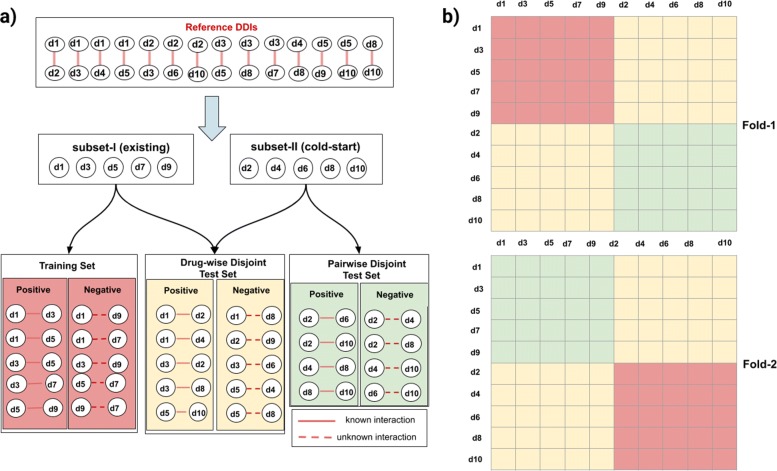
Fig. 4.
To illustrate a partitioning for 2-fold cross validation, we consider a toy example of DDI prediction, in which the reference data has 10 drugs and 14 DDIs. a The train-test generation workflow for a fold. Drugs are randomly split into 2 groups where one group is used as cold-start drugs to generate test sets, while the remaining drugs (existing) are used to generate a training set for a fold. Splitting the drugs into 2 groups leads to partitioning of drug pairs into 3 sets: training set, drug-wise disjoint test set and pairwise disjoint test set. The pairs which include both components from existing drugs are assigned to the training set. The interactions between existing drugs and cold-start drugs are assigned to the drug-wise disjoint test set and likewise, the interactions between cold-start drugs are assigned to the pairwise disjoint test set. In other words, the drug-wise test set will contain the drug pairs each of which shares only one element with training set. The pairwise test set will contain the drug pairs in which neither component is shared with the training set. b Partitioning of the drug-drug pair space into training and test subsets for each fold. The pair space is represented by a table with 10×10 cells. The drug-drug pair space is divided into different blocks, which account for training, drug-wise testing and pairwise testing parts, and are filled with red, yellow and green colors respectively