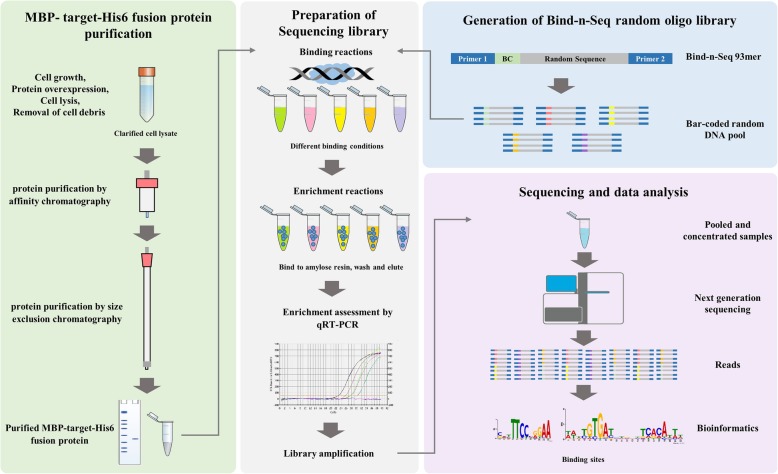
Fig. 1.
Bind-n-seq experimental overview. The protein purification strategy depends on the properties of the target protein and should be optimized in each case. For YipR, both MBP and His affinity tags were incorporated and an affinity chromatography step was followed by a size exclusion step. After purification, the target protein is assessed for concentration, stability and purity. The protein quality is an essential requirement (green panel left). The Bind-n-seq substrate is an oligo containing constant regions (Primer A and Primer B) a 3-nucleotide bar code (BC) and 21 bp random region (blue panel right). Barcoded oligonucleotides are mixed with various proteins, washed to remove unbound DNA, pooled and sequenced with short read technology (grey panel middle). Reads are sorted by their bar codes and processed through several bioinformatics procedures that result in motifs corresponding to the DNA binding sites of each protein (pink panel right)