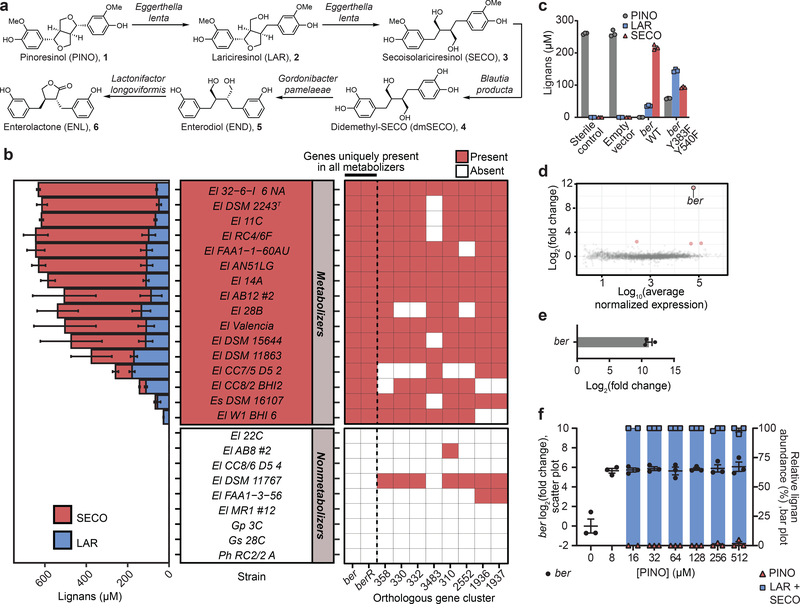
Figure 1. Lignan metabolism varies between Coriobacteriia strains and enables identification of a single enzyme sufficient to catalyze the first two reactions in the lignan metabolism pathway.
a, Known biotransformations by which gut-residing bacteria can convert pinoresinol (PINO) into the enterolignan products enterodiol (END) and enterolactone (ENL). b, Evaluation of PINO-metabolizing ability across our collection of strains from the Coriobacteriia class, of which E. lenta is a member. Lignans were quantified by HPLC (values are mean±SEM, n=3 biologically independent samples). ber: benzyl ether reductase; berR: benzyl ether reductase regulator; El: Eggerthella lenta; Es: Eggerthella sinesis; Gp: Gordonibacter pamelaeae; Gs: Gordonibacter species; Ph: Paraeggerthella hongkongensis. c, Incubation of PINO (250 μM) with E. coli Rosetta 2(DE3) expressing WT or mutagenized Ber on a pET19btev plasmid. PINO and the products of its benzyl ether reduction, laricirisinol (LAR) and secoisolaricerisinol (SECO), were quantified by HPLC (bars are mean±SEM, n=3 biologically independent samples). d, RNAseq results obtained upon exposure of E. lenta DSM2243T to PINO (n=3 biologically independent samples) relative to vehicle (n=3 biologically independent samples). Points in pink are genes for which was observed a fold-change>|4| and FDR<0.01 (Wald test, Benjamini–Hochberg multiple testing correction). e, RT-qPCR confirmation of ber up-regulation upon exposure of E. lenta DSM2243T to PINO (500 μM) relative to vehicle (bar is mean±SEM, n=3) and f, at varying concentrations of PINO (mean±SEM, n=3 biologically independent samples), along with conversions of PINO to LAR and SECO at respective doses of PINO (bar is mean±SEM, n=3 biologically independent samples; except 16 μM, n=2 biologically independent samples). Lignan concentrations measured by HPLC. Lignan conversion data not available for 8 μM of PINO due to instrument’s limit of quantitation.