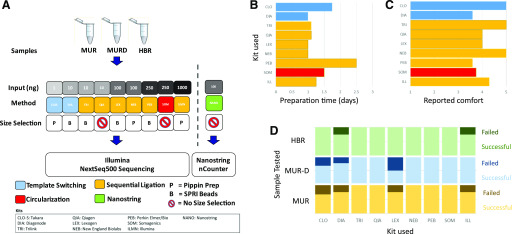
FIGURE 1.
Study design and workflow metrics. A) Schematic of study design is shown. MUR, MUR-D, and HBR were processed using 9 different smRNA profiling methods at 4 sites each. The general methodologies included sequential ligation (Illumina, TriLink, Qiagen, NEB, PerkinElmer, Lexogen), template switching (Takara, Diagenode), circularization (Somagenics), and NanoString probe-based hybridization. B) Total start-to-finish preparation time for each kit as reported by sites. C) Mean “ease of use” reported by each site (scale: 1 = uncomfortable, 5 = comfortable). D) Library preparation success rates. Light color blocks are successfully produced libraries. Dark colors are failed libraries. CLO and CLO-S, Takara Bio (Clontech) SMARTer smRNA-Seq Kit; DIA, Diagenode CATS Small RNA-Seq Kit; ILL and ILMN, Illumina TruSeq Small RNA Library Prep Kit; LEX, Lexogen Small RNA-Seq Library Prep Kit; NANO, NanoString nCounter miRNA Expression Assay; NEB, New England Biolabs NEBNext Small RNA Library Prep Set; PEB, PerkinElmer NextFlex Small RNA-Seq Kit v.3; QIA, Qiagen QIAseq miRNA Library Kit; SOM, Somagenics RealSeq-AC miRNA Library Kit; TRI, Trilink CleanTag Small RNA Library Prep Kit.