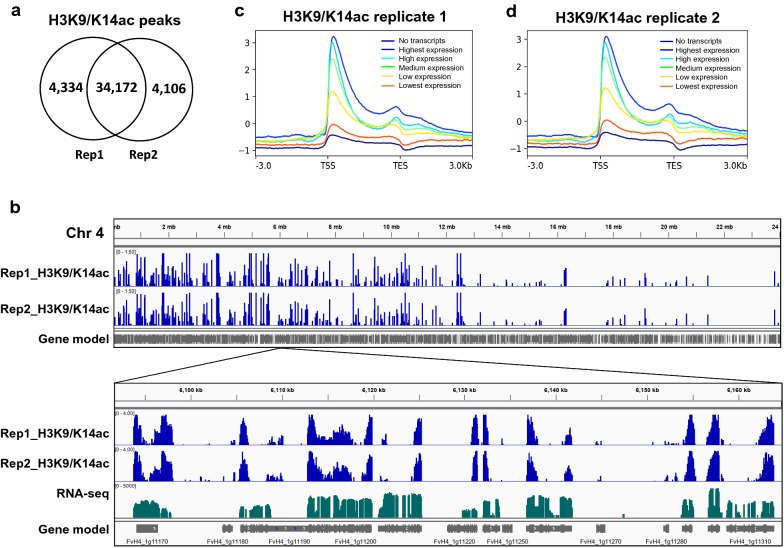
Fig. 4.
The consistency and reproducibility of our N-ChIP protocol validated by ChIP-seq profiles. a A venn diagram showing the overlap between the H3K9/K14ac-enriched peaks called by ChIP-seq replicate 1 and 2. b IGV browser shots illustrating chromosome-wide and local H3K9/K14ac enrichments for the two biological replicates. c, d Metagene analysis illustrating a positive correlation between local H3K9/K14ac enrichment and gene expression levels. The expressed genes were evenly classified into five groups according to their transcript levels (blue-highest expression, light blue-high, green-medium, yellow-low and red-lowest). The group of genes with no detectable transcripts was included as well (black line). The transcriptome data were published previously [30]