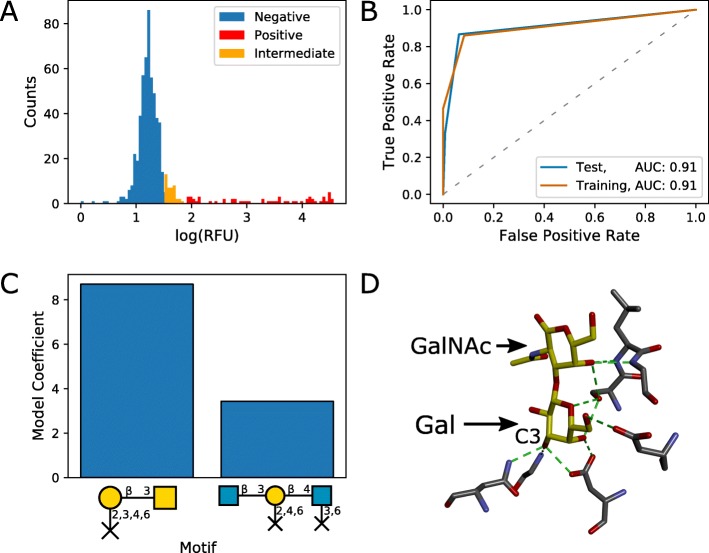
Fig. 3.
Predicted carbohydrate-binding motifs of PNA from CFG glycan microarray data. a Distribution of RFUs and classification of non-binding (blue), intermediate binding (orange), and binding glycans (red). b ROC curves for the test (n=143) and training (n=428) sets. The ratio of negative to positive samples was 9.0. c Logistic regression coefficients for identified motifs. d The intermolecular hydrogen bonding interactions (shown in green) between the T antigen (carbon backbone shown in yellow) and the carbohydrate-binding domain of peanut agglutinin (PNA) (carbon backbones shown in grey). Carbon 3 of the Gal monomer is labelled to indicate where the sialic acid is linked in the sialyl T antigen. Reproduced from an X-ray crystal structure at 2.5 Å resolution available at the PDB (PDB: 2TEP) [28]. See Additional file 1 for a detailed notation key